SciScholar
Health Warn
- License — License: MIT
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Low visibility — Only 9 GitHub stars
Code Pass
- Code scan — Scanned 12 files during light audit, no dangerous patterns found
Permissions Pass
- Permissions — No dangerous permissions requested
This project provides an open-source Python client for interacting with the SciNet API. It is designed to help automate scientific research tasks, such as generating and evaluating academic ideas, predicting research trends, and profiling authors using a massive academic knowledge graph.
Security Assessment
Overall Risk: Low. The tool functions as an API client and makes external network requests to a hosted SciNet server to retrieve data. The automated code scan of 12 files found no dangerous patterns, no hardcoded secrets, and no risky shell command executions. Users should simply be aware that the inputs and generated research requests are sent externally to the third-party API.
Quality Assessment
The project is actively maintained, with its last push occurring today. It is released under the permissive and standard MIT license, making it highly accessible for developers. However, it currently has extremely low community visibility with only 7 GitHub stars, indicating it is likely a new or niche academic tool without a broad established user base to validate it.
Verdict
Safe to use, though developers should keep in mind that all task executions and queries are processed via an external hosted API.
A Large-Scale Knowledge Graph for Automated Scientific Research
SciScholar: A Large-Scale Knowledge Graph for Automated Scientific Research
🌐 English · 简体中文
A pip-installable client and CLI for literature-grounded scientific research workflows on top of the hosted SciNet API.
📄 arXiv · 🔑 Get API Token · 🩺 API Health
✨ Overview
SciNet is a research map you can use from the command line. Give it a topic, an idea, an author, or a paper trail, and it helps you look up literature, gather graph-backed evidence, and turn the result into readable reports and reusable JSON artifacts.
Behind that simple workflow is a large scientific knowledge graph. SciNet connects papers, authors, institutions, venues, keywords, citations, and a four-level research taxonomy from domains down to topics. That means a search is not limited to matching words: it can follow how research areas, people, concepts, and papers relate to one another.
This repository packages that capability as a lightweight SciNet client. New users can install it with pip, register an API token, and start running literature-grounded research tasks without setting up Neo4j, maintaining graph data, or touching backend infrastructure.
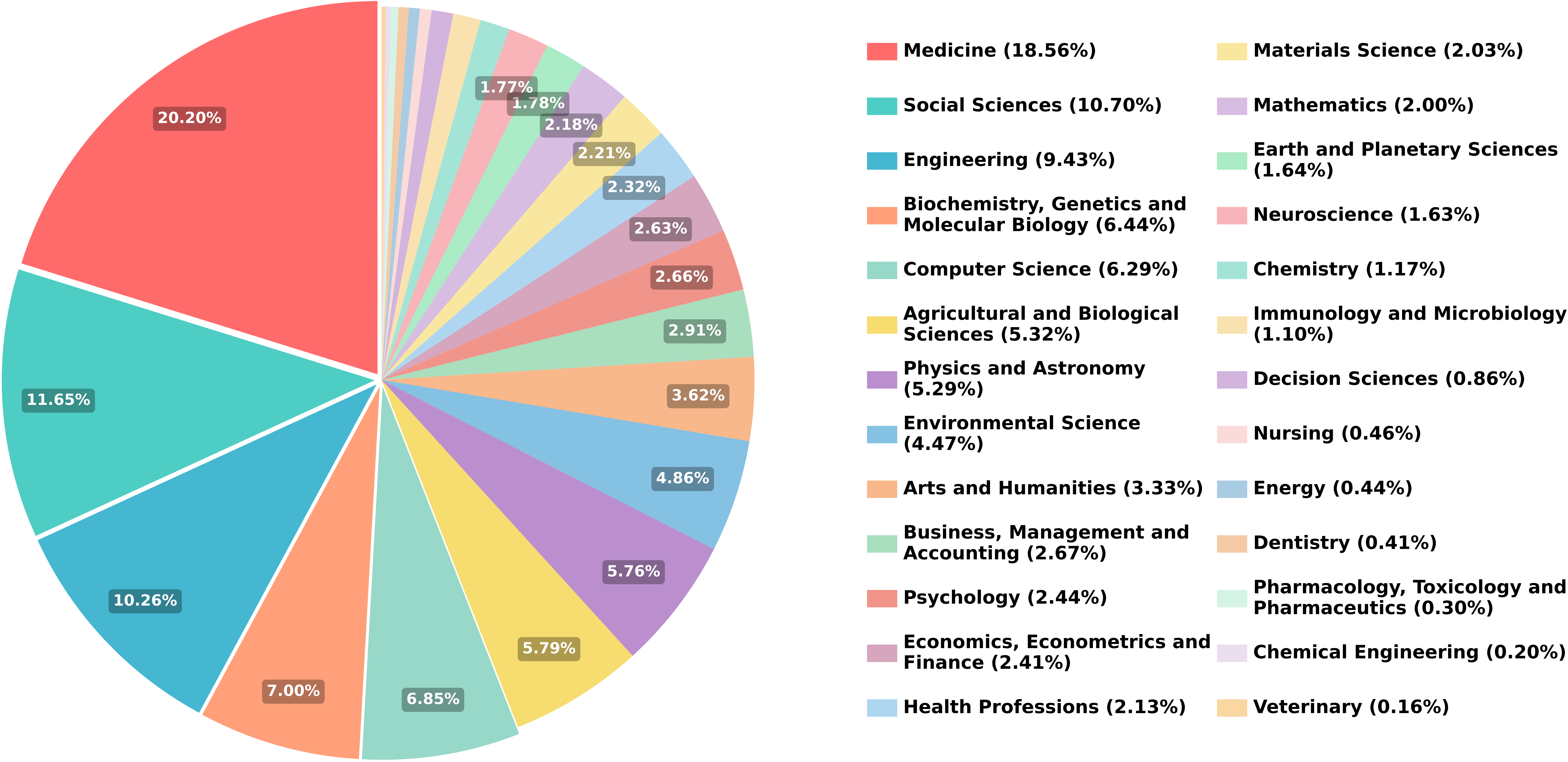
SciNet spans a broad research landscape, from medicine and social sciences to engineering, computer science, materials science, mathematics, and more.
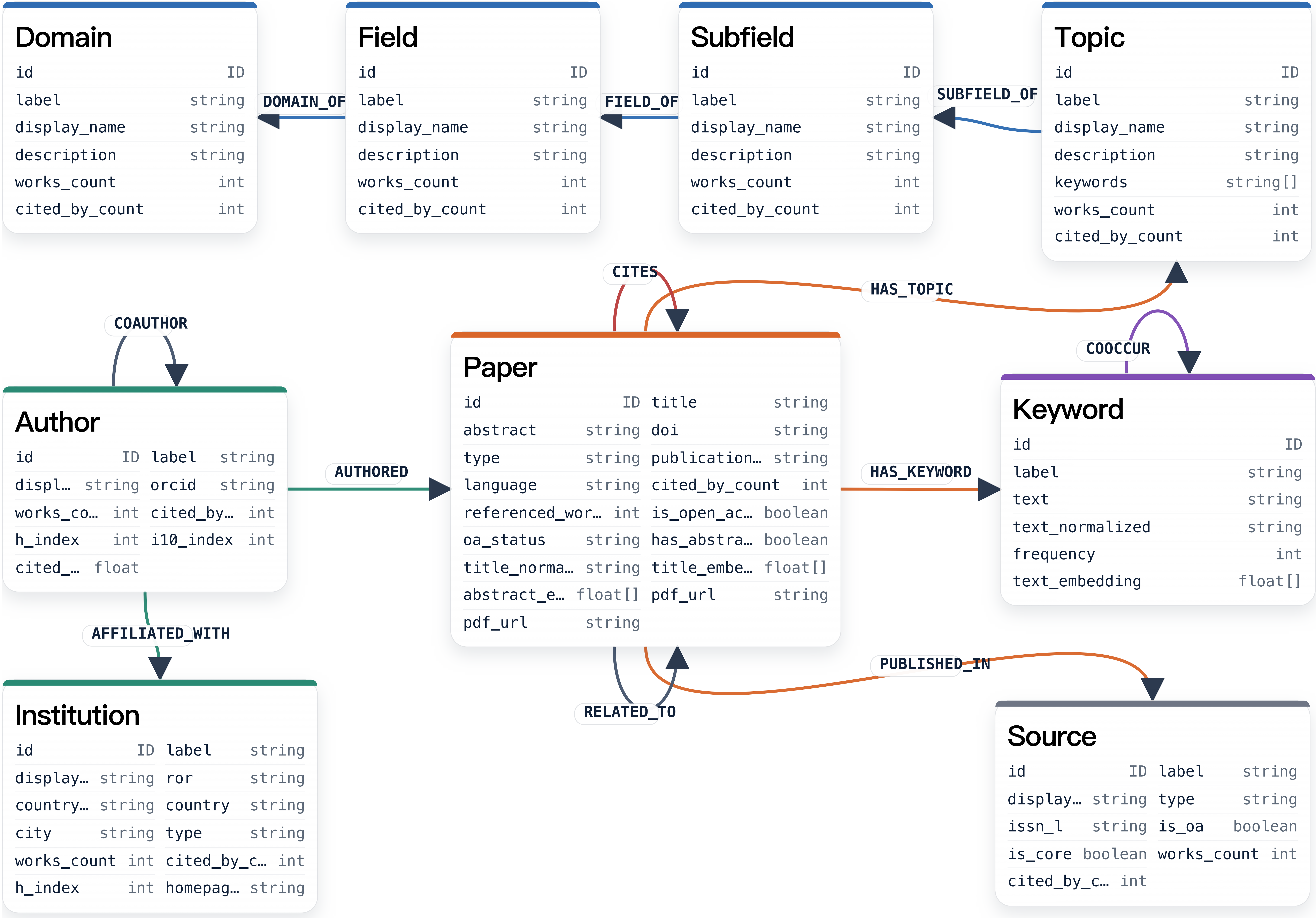
The graph links papers with authors, institutions, sources, keywords, citations, related work, and the domain-field-subfield-topic hierarchy.
With the client, SciNet becomes a practical research assistant for:
- graph-aware paper search: combine keywords, semantic matching, title anchors, references, and graph propagation instead of stopping at plain keyword matching;
- research workflow automation: run literature review, idea grounding, idea evaluation, idea generation, trend analysis, related-author retrieval, and researcher profiling;
- agent-friendly outputs: keep reproducible machine-readable artifacts such as
request.jsonandresponse.json, plus user-facingsummary.txtandreport.md; - editable CLI skills: inspect, copy, modify, and rerun common downstream workflows as reusable JSON skills;
- portable Agent Skill pack: use the packaged skills in
agent-skill/to let tools such as Codex, Claude Code, and other coding agents invoke SciNet workflows with the right defaults and artifact-reading habits.
📑 Table of Contents
- ✨ Overview
- 📑 Table of Contents
- 🚀 Quick Start
- 🔑 API Token
- 🧩 Supported Tasks
- 🛠️ CLI-First Workflow
- 🧰 Editable Skills
- Agent Skill
- 🐍 Python SDK
- 📦 Outputs and Artifacts
- 📂 Repository Layout
- 🧯 Troubleshooting
- 📝 TODO
- ✍️ Citation
- 📄 License
🚀 Quick Start
1. Install
Install directly from GitHub:
pip install "git+https://github.com/zjunlp/SciNet.git#subdirectory=scinet"
For isolated CLI usage:
pipx install "git+https://github.com/zjunlp/SciNet.git#subdirectory=scinet"
After installation:
scinet -h
2. Register an API Token
Open:
http://scinet.openkg.cn/register
Complete email verification and copy your personal token.
Quick link: 🔑 API Token.
3. Configure
At minimum, configure the hosted SciNet API endpoint and your personal token.
Linux / macOS:
export SCINET_API_BASE_URL="http://scinet.openkg.cn"
export SCINET_API_KEY="your-personal-scinet-token"
export SCINET_TIMEOUT=900
export SCINET_RUNS_DIR="./runs"
Windows CMD:
set SCINET_API_BASE_URL=http://scinet.openkg.cn
set SCINET_API_KEY=your-personal-scinet-token
set SCINET_TIMEOUT=900
set SCINET_RUNS_DIR=.\runs
Compatibility variables:
KG2API_BASE_URL=http://scinet.openkg.cn
KG2API_API_KEY=your-personal-scinet-token
For new setups, prefer SCINET_*.
📕 Optional: use your own LLM for keyword extraction
export LLM_PROVIDER="chat_completions"
export LLM_API_KEY="your-provider-api-key"
export LLM_BASE_URL="https://your-provider-or-gateway.example/v1"
export LLM_MODEL="your-model-name"
# Optional when your provider uses a custom endpoint or auth header:
# export LLM_CHAT_COMPLETIONS_URL="https://your-provider-or-gateway.example/v1/chat/completions"
# export LLM_AUTH_HEADER="x-api-key: your-provider-api-key"
export SCINET_LLM_TIMEOUT=30
export SCINET_LLM_TEMPERATURE=0
export SCINET_LLM_MAX_TOKENS=512
This step is optional. Configure it only when you want SciNet to use your LLM API to turn a free-form query into better search keywords.
Keep LLM_PROVIDER=chat_completions, then replace LLM_API_KEY, LLM_BASE_URL, and LLM_MODEL with your provider values. If your provider gives a full chat-completions endpoint, set LLM_CHAT_COMPLETIONS_URL; if it requires a custom auth header, set LLM_AUTH_HEADER.
Leave the LLM values empty if you do not need this. SciNet will use built-in keyword extraction, and normal search, review, idea, trend, and researcher workflows still run.
User-editable template: .env.example. Set these variables only if you want LLM-assisted keyword extraction.
🖊 Optional: OpenAlex metadata support
export OA_API_KEY=""
export OPENALEX_MAILTO=""
OpenAlex is useful when you want extra metadata or PDF-related support. It is not required for the main CLI examples in this README. If you leave these variables empty, normal SciNet retrieval still works.
User-editable template: .env.example. Set these only if you want OpenAlex-assisted metadata support.
🖌 Optional: GROBID for local PDF workflows
GROBID is only needed when you process local PDF files. It reads scientific PDFs and extracts titles, authors, abstracts, and references. If you are only running the text-based CLI commands above, you can skip this section.
Start GROBID locally:
docker pull lfoppiano/grobid:latest
docker run -d --rm --name grobid -p 8070:8070 lfoppiano/grobid:latest
curl http://127.0.0.1:8070/api/isalive
Then set:
export GROBID_BASE_URL="http://127.0.0.1:8070"
Windows CMD:
set GROBID_BASE_URL=http://127.0.0.1:8070
User-editable template: .env.example. Leave GROBID_BASE_URL empty unless you process local PDFs.
Runtime variables:
| Variable | Required For | Notes |
|---|---|---|
SCINET_API_BASE_URL |
all hosted SciNet tasks | Hosted SciNet API base URL. |
SCINET_API_KEY |
all hosted SciNet tasks | Sent as X-API-Key and Authorization: Bearer. |
LLM_PROVIDER |
optional frontend enhancement | Keep as chat_completions. |
LLM_API_KEY |
optional frontend enhancement | Your provider key; leave empty for local or no-auth services. |
LLM_BASE_URL |
optional frontend enhancement | Provider base URL, usually ending in /v1. |
LLM_CHAT_COMPLETIONS_URL |
optional frontend enhancement | Use only when your provider gives a full endpoint. |
LLM_MODEL |
optional frontend enhancement | Model name from your provider. |
LLM_AUTH_HEADER |
optional frontend enhancement | Use only for custom auth, for example x-api-key: your-provider-api-key. |
LLM_HTTP_HEADERS |
optional frontend enhancement | Optional extra headers as JSON. |
GROBID_BASE_URL |
PDF tasks | Needed for --pdf-path workflows. |
OA_API_KEY |
optional | OpenAlex metadata/PDF support. |
OPENALEX_MAILTO |
optional | OpenAlex contact email. |
4. Test
scinet health
scinet config
5. Run a Paper Search
scinet search-papers \
--query "open world agent" \
--keyword "high:open world agent" \
--top-k 10
🔑 API Token
SciNet uses personal API tokens for public access.
Browser Registration
Visit:
http://scinet.openkg.cn/register
Steps:
- enter your name, email, organization, and use case;
- click Send code;
- check your inbox for the verification code;
- enter the code and create a token;
- copy the returned
scinet_xxxtoken.
The token is shown only once.
Check Token Status
curl -H "Authorization: Bearer $SCINET_API_KEY" \
http://scinet.openkg.cn/v1/auth/token/status
Check Usage
curl -H "Authorization: Bearer $SCINET_API_KEY" \
"http://scinet.openkg.cn/v1/auth/usage?days=7"
🧩 Supported Tasks
| Command | Scenario | Main Output |
|---|---|---|
scinet search-papers |
Paper search | Related papers and Markdown report |
scinet related-authors |
Related-author discovery | Candidate authors and scores |
scinet author-papers |
Author paper lookup | Papers by a specified author |
scinet support-papers |
Support-paper retrieval | Evidence papers for candidate authors |
scinet paper-search |
Lightweight low-level paper search | Fast paper candidates |
scinet literature-review |
Literature review | Core paper pool, timeline, writing hints |
scinet idea-grounding |
Idea grounding | Similar works and differentiation evidence |
scinet idea-evaluate |
Idea evaluation | Evidence for novelty, feasibility, and soundness |
scinet idea-generate |
Idea generation | Topic combinations and idea seeds |
scinet trend-report |
Trend analysis | Evolution evidence and representative works |
scinet researcher-review |
Researcher background review | Research trajectory and representative works |
scinet skill |
Editable skill registry | Reusable workflow presets |
🛠️ CLI-First Workflow
SciNet is CLI-first: you can start with one command, inspect the saved artifacts, and then move into larger research workflows. If you are new, run help once, try a basic retrieval, then choose one of the downstream workflows below.
Documentation: 📚 SciNet Documentation. Use it to check API setup, CLI commands, parameter meanings, and runnable examples.
Help
scinet -h
scinet search-papers -h
scinet literature-review -h
scinet skill -h
Input Styles
SciNet supports two input styles. For formal runs, prefer expert parameters because every field is explicit and easier to reproduce. Natural-language input is useful for quick trials or exploratory use.
Recommended: expert parameters
scinet --timeout 900 search-papers \
--retrieval-mode hybrid \
--query "open world agent" \
--domain "artificial intelligence" \
--time-range 2020-2024 \
--keyword "high:open world agent" \
--keyword "middle:embodied agent" \
--title "middle:Voyager: An Open-Ended Embodied Agent with Large Language Models" \
--reference "low:JARVIS-1: Open-World Multi-task Agents with Memory-Augmented Multimodal Language Models" \
--top-k 5 \
--top-keywords 0 \
--max-titles 0 \
--max-refs 0 \
--bias-keyword high \
--bias-related high \
--bias-exploration low \
--ranking-profile precision \
--report-max-items 5
Compatible: natural-language input
Use --text when you want SciNet to parse the request from a short instruction. You can still add structured hints such as keyword[high]: ... in the text.
scinet --timeout 900 search-papers \
--retrieval-mode hybrid \
--text "Find papers related to open world agent in artificial intelligence since 2020. Return 3 papers.
keyword[high]: open world agent" \
--top-k 3 \
--top-keywords 1 \
--max-titles 0 \
--max-refs 0
Basic Retrieval
Use this when you want a quick, evidence-backed paper list for one topic.
scinet search-papers \
--query "open world agent" \
--domain "artificial intelligence" \
--time-range 2020-2024 \
--keyword "high:open world agent" \
--top-k 5 \
--top-keywords 0 \
--max-titles 0 \
--max-refs 0
Downstream Workflows
Each workflow prints a concise terminal summary and saves full artifacts under runs/<run_id>/.
Literature Review
Build an initial reading list and get evidence for writing a literature review.
scinet literature-review \
--query "retrieval augmented generation" \
--domain "artificial intelligence" \
--time-range 2020-2025 \
--keyword "high:retrieval augmented generation" \
--top-k 10
Idea Evaluation
Check whether a proposed research idea is novel, feasible, and well supported by existing work.
scinet idea-evaluate \
--idea "LLM-based multi-perspective evaluation for scientific research ideas" \
--domain "artificial intelligence" \
--time-range 2020-2025 \
--keyword "high:idea evaluation" \
--keyword "middle:LLM as a judge" \
--top-k 10
Idea Generation
Explore promising topic combinations and generate candidate research directions.
scinet idea-generate \
--query "knowledge editing for large language models" \
--domain "artificial intelligence" \
--time-range 2020-2025 \
--keyword "high:knowledge editing" \
--keyword "middle:large language models" \
--keyword "low:continual learning" \
--top-k 10
Trend Report
Trace how a topic has developed and identify representative works along the way.
scinet trend-report \
--query "retrieval augmented generation" \
--domain "artificial intelligence" \
--time-range 2020-2025 \
--keyword "high:retrieval augmented generation" \
--keyword "middle:knowledge graph" \
--top-k 10
Researcher Review
Summarize a researcher's publication trajectory and representative papers.
scinet researcher-review \
--author "Yoshua Bengio" \
--limit 10 \
--no-abstract
Retrieval Modes
| Mode | Meaning | Best For |
|---|---|---|
keyword |
Keyword-driven KG retrieval | Clear terminology |
semantic |
Semantic retrieval | Broad semantic matching |
title |
Title-anchor retrieval | Known paper titles |
hybrid |
Keyword + semantic + title + graph walk | Default and recommended |
If --retrieval-mode is omitted, SciNet uses hybrid.
Expert Anchors
Use anchors when you already know a strong keyword, title, or reference and want the graph search to start from it.
--keyword "high:open world agent"
--title "middle:Voyager: An Open-Ended Embodied Agent with Large Language Models"
--reference "low:JARVIS-1: Open-World Multi-task Agents with Memory-Augmented Multimodal Language Models"
Graph Bias Parameters
| Parameter | Meaning |
|---|---|
--bias-keyword |
Keyword association strength |
--bias-non-seed-keyword |
Non-seed keyword expansion |
--bias-citation |
Citation edge strength |
--bias-related |
Paper relatedness strength |
--bias-authorship |
Author-paper relation strength |
--bias-coauthorship |
Coauthor network strength |
--bias-cooccurrence |
Keyword co-occurrence strength |
--bias-exploration |
Graph exploration level |
--ranking-profile |
Ranking preference: precision, balanced, discovery, impact |
Recommended safe defaults:
--top-k 10
--top-keywords 0
--max-titles 0
--max-refs 0
--bias-exploration low
🧰 Editable Skills
SciNet skills are JSON presets for downstream research workflows. They make complex workflows easier to inspect, reuse, and customize.
scinet skill list
scinet skill show literature-review
scinet skill run literature-review --query "open world agent" --keyword "high:open world agent"
scinet skill run --dry-run literature-review --query "open world agent" --keyword "high:open world agent"
Create a custom skill:
scinet skill init my-review --from literature-review
This creates:
./skills/my-review.json
Edit it, then run:
scinet skill run my-review --query "your topic"
User-defined skills are loaded from:
./skills/*.json~/.scinet/skills/*.json- directories specified by
SCINET_SKILLS_DIR
User-defined skills can override built-in skills with the same name.
Agent Skill
SciNet also ships a portable Agent Skill pack under agent-skill/. These are not runtime outputs; they are reusable skill packs that teach tools such as Codex, Claude Code, and other coding agents how to choose and run SciNet workflows, pass reliable parameters, and read saved artifacts.
Included skills:
| Skill | Workflow | Use case |
|---|---|---|
scinet-literature-review |
literature-review |
Reading lists and related-work reports |
scinet-idea-grounding |
idea-grounding |
Prior-art grounding for research ideas |
scinet-idea-evaluate |
idea-evaluate |
Novelty, feasibility, and soundness checks |
scinet-idea-generate |
idea-generate |
Literature-grounded idea seeds |
scinet-trend-report |
trend-report |
Timeline and trend analysis |
scinet-researcher-review |
researcher-review |
Researcher profiles and representative works |
scinet-quick-paper-search |
paper-search |
Fast paper candidate lookup |
To use one locally, copy its directory into the skill directory supported by your agent tool, then restart or refresh that tool. For Codex, that is usually ~/.codex/skills or %USERPROFILE%\.codex\skills. The CLI presets remain in scinet/src/scinet/builtin_skills.json; the Agent Skill pack is the agent-facing layer on top.
🐍 Python SDK
SciNet also provides a lightweight Python client.
from scinet import SciNetClient
client = SciNetClient()
print(client.health())
result = client.search_papers(
query="open world agent",
keywords=[{"text": "open world agent", "score": 10}],
top_k=3,
)
print(result)
You can also pass credentials directly:
from scinet import SciNetClient
client = SciNetClient(
base_url="http://scinet.openkg.cn",
api_key="your-personal-scinet-token",
)
print(client.token_status())
📦 Outputs and Artifacts
Terminal output is concise and table-based. Full outputs are saved under:
runs/<run_id>/
Common artifacts:
| File | Description |
|---|---|
plan.json |
Structured search plan |
request.json |
Full request sent to SciNet API |
response.json |
Raw backend response |
summary.txt |
Short summary |
report.md |
User-facing Markdown report |
metadata.json |
Runtime metadata |
📂 Repository Layout
The tree below highlights the main user-facing areas of the repository. Generated outputs and local cache folders are omitted.
SciNet/
README.md / README_zh.md # project documentation
.env.example # root runtime configuration template
requirements.txt
run_scinet.py # lightweight local runner
agent-skill/ # portable Agent Skill pack
docs/api/ # unified static API and CLI documentation site
imgs/ # README figures
scinet/ # pip-installable SciNet client package
pyproject.toml
src/scinet/ # packaged CLI, client, config, and skills
core/ search/ tasks/ # retrieval planning and workflow logic
evidence/ llm/ renderers/ # PDF evidence, optional LLM, report rendering
examples/ tests/
references/search/ # reference KG search implementation
runs/ # generated CLI outputs
🧯 Troubleshooting
scinet health works but search-papers returns 401
Your token is missing or invalid.
echo $SCINET_API_KEY
export SCINET_API_KEY="your-personal-scinet-token"
Windows CMD:
set SCINET_API_KEY=your-personal-scinet-token
No email verification code
Check the email address, spam folder, and resend interval.
Retrieval is slow or times out
Use lightweight settings:
--top-k 3
--top-keywords 0
--max-titles 0
--max-refs 0
--bias-exploration low
scinet command is not found on Windows
Use the virtual environment executable directly:
.venv\Scripts\scinet.exe -h
or reinstall:
.venv\Scripts\python.exe -m pip install -e .
📝 TODO
- CLI Tools. Add more user-facing CLI capabilities so downstream users and AI agents can invoke retrieval workflows without touching database internals.
- Portable Agent Skill pack. Package reusable agent skills for common scientific discovery workflows and expose best practices as easier-to-load components.
- More Knowledge. Integrate more knowledge forms beyond paper-centric entities, such as datasets, code, standards, theorems, and experimental experience.
- Benchmark and Evaluation. Build dedicated benchmarks and evaluation protocols for downstream scientific research tasks supported by SciNet.
- Dynamic Update Improve dynamic knowledge updates toward a more systematic and frequent refresh mechanism.
✍️ Citation
If you find SciNet helpful, please cite:
📄 License
This project is licensed under the MIT License. See LICENSE for details.
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found


