Genomic-Agent-Discovery
Health Warn
- License — License: MIT
- Description — Repository has a description
- Active repo — Last push 0 days ago
- Low visibility — Only 5 GitHub stars
Code Warn
- network request — Outbound network request in dashboard-react/src/App.jsx
- network request — Outbound network request in dashboard-react/src/SetupScreen.jsx
Permissions Pass
- Permissions — No dangerous permissions requested
This tool is an MCP server that analyzes raw DNA files (from services like 23andMe or AncestryDNA) using a collaborative team of AI agents to generate a comprehensive health report.
Security Assessment
Overall Risk: Medium. This tool processes highly sensitive personal data—your raw genetic information. While the documentation explicitly claims that everything runs locally and "nothing is uploaded anywhere," the static scan flagged outbound network requests in the frontend dashboard files (`App.jsx` and `SetupScreen.jsx`). These requests are likely required to connect to LLM providers (like Anthropic or OpenAI) to run the AI agents, meaning your queries or snippets of data may leave your machine depending on how you configure the application. No hardcoded secrets or dangerous background shell permissions were detected, but users should be acutely aware that local DNA processing requires trusting the code's execution path and the external LLM API endpoints it communicates with.
Quality Assessment
The project is very new and currently has low visibility with only 5 GitHub stars, meaning it has not been extensively vetted by the broader developer community. However, it uses a standard permissive MIT license, includes a clear description, and shows signs of active development with very recent repository updates.
Verdict
Use with caution. The tool operates exactly as intended for local genetic analysis, but given the extreme sensitivity of genomic data, it should only be used if you fully audit the code and strictly manage your external LLM API routing.
Drop in your 23andMe/AncestryDNA file. Watch AI agents discuss your genome in real-time.
Genomic Agent Discovery
AI agents that collaborate to analyze your DNA. Open source. Runs locally. Your data never leaves your machine.
Quick Start • Dashboard • Presets • Agent Prompts • Architecture • Configuration • Database • Privacy





Upload your raw DNA file from 23andMe, AncestryDNA, MyHeritage, FamilyTreeDNA, or any VCF -- and watch a team of AI agents fan out across 12+ public genomics databases, share discoveries with each other in real time, and produce a comprehensive health report. Everything runs on your machine. Nothing is uploaded anywhere.
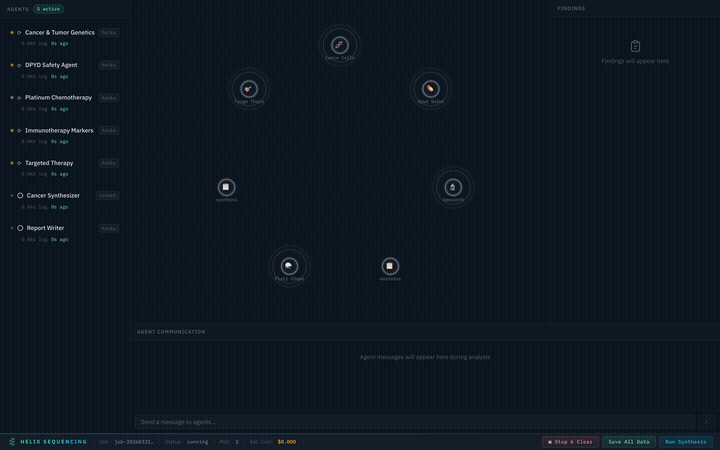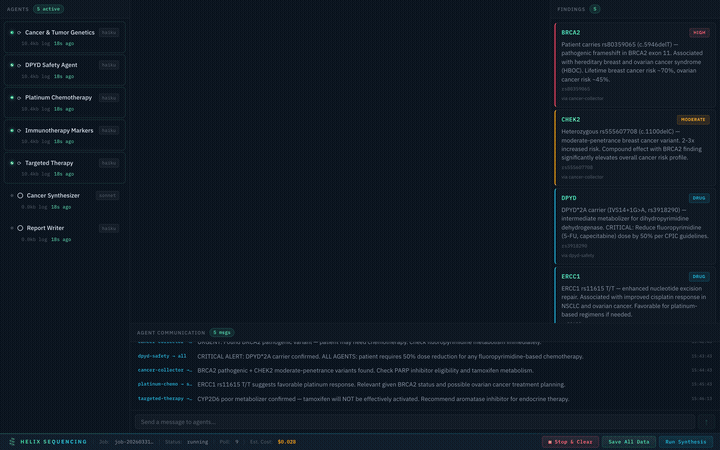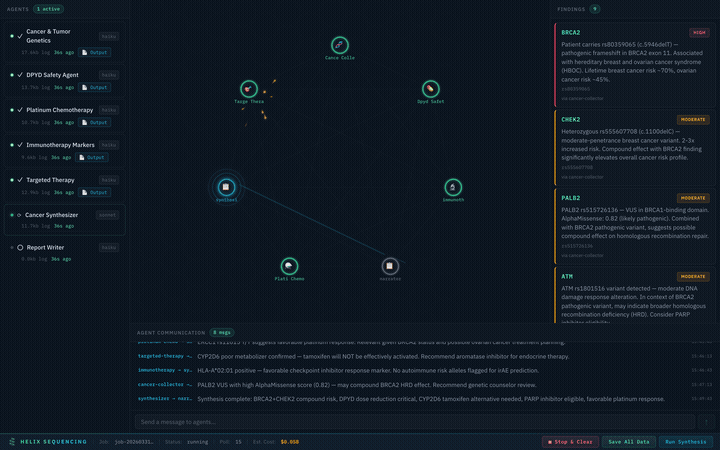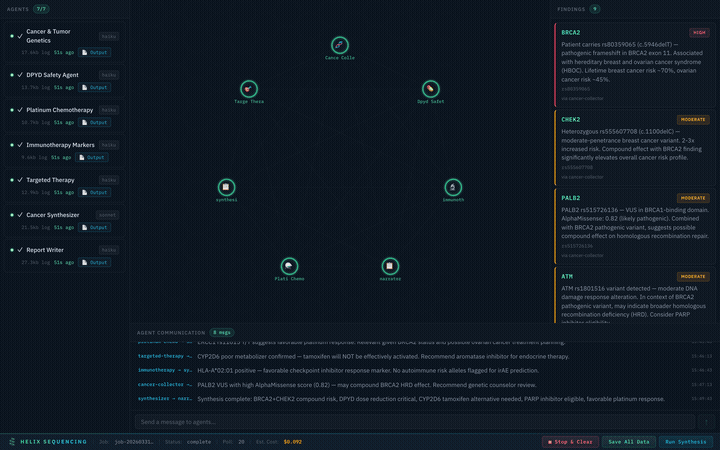
Real-time pipeline: 7 agents collaborating on a cancer genomics analysis
Quick Start
Have a Claude Pro or Max subscription? (Recommended)
No API key needed. Your subscription covers everything.
# 1. Install Claude CLI if you haven't already
npm install -g @anthropic-ai/claude-code
# 2. Log in once — opens browser OAuth (free, uses your Claude Pro/Max subscription)
claude login
# 3. Clone, build, and run
git clone https://github.com/HelixGenomics/Genomic-Agent-Discovery.git
cd Genomic-Agent-Discovery
npm install && npm run build-db
npm start -- --dna ~/Downloads/my-dna-raw.txt
A dashboard opens in your browser and you can watch the agents work. That's it — no API keys, no per-token charges.
Using an Anthropic API key instead
export ANTHROPIC_API_KEY=sk-ant-... # get one at console.anthropic.com
git clone https://github.com/HelixGenomics/Genomic-Agent-Discovery.git
cd Genomic-Agent-Discovery
npm install && npm run build-db
npm start -- --dna ~/Downloads/my-dna-raw.txt --provider anthropic-api
Typical cost: $1–5 per analysis run depending on preset. See Provider options for OpenAI, Gemini, Ollama, and others.
Dashboard
The dashboard is a real-time mission control for your genomic analysis. It provides full visibility into agent status, findings, inter-agent communication, and costs — all in a single page.
Setup Panel
When you launch the dashboard, you'll see the setup panel where you configure your analysis before starting.
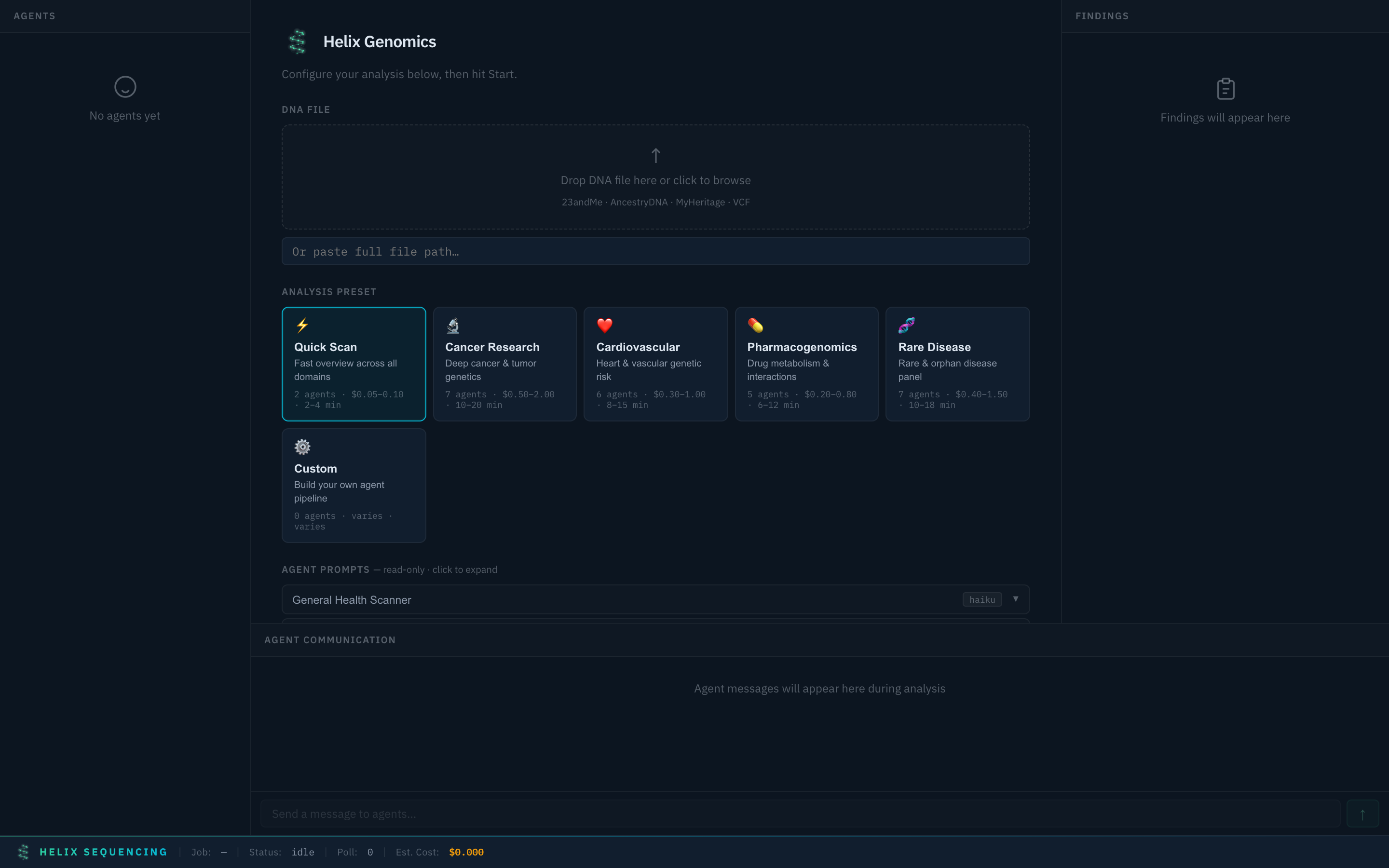
Setup panel — select a preset, configure settings, and start your analysis
Preset Selection
Choose from 6 built-in research presets, each tuned for a specific domain. Selecting a preset instantly configures the agent pipeline, prompts, models, and focus areas.
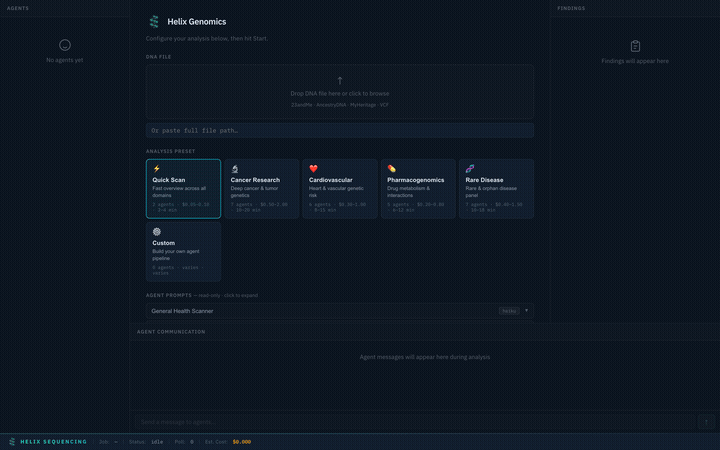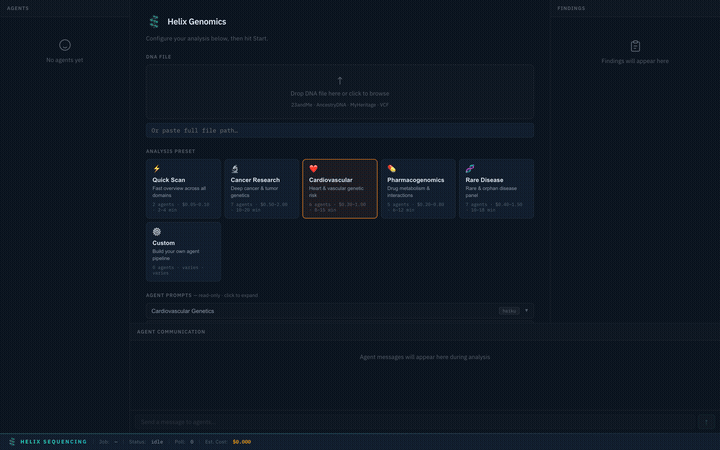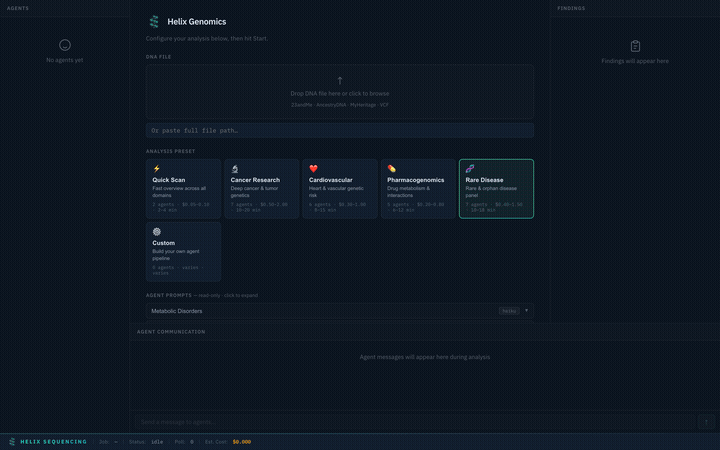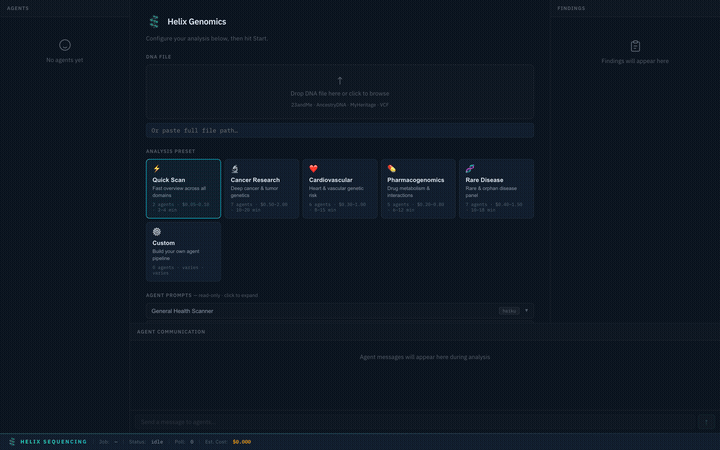
Switch between presets — each configures a different agent team with specialized prompts
Available presets:
| Preset | Agents | Est. Cost | Focus |
|---|---|---|---|
| Quick Scan ⚡ | 2 | $0.05–0.10 | Fast overview across all domains |
| Cancer Research 🔬 | 7 | $0.50–2.00 | Deep cancer & tumor genetics with DPYD safety, platinum chemo, immunotherapy, and targeted therapy agents |
| Cardiovascular ❤️ | 6 | $0.30–1.00 | Lipid genetics, arrhythmia risk, coagulation, and structural heart |
| Pharmacogenomics 💊 | 4 | $0.20–0.80 | CYP enzyme panel, drug transporters, and full CPIC pharmacogene coverage |
| Rare Disease 🧬 | 7 | $0.40–1.50 | Metabolic disorders, neurological conditions, connective tissue, immunodeficiency, and rare cancer syndromes |
| Custom ⚙️ | You decide | Varies | Build your own agent pipeline from scratch |
Agent Prompts
Every preset comes with full, detailed prompts visible in an expandable accordion. You can review exactly what each agent will investigate before starting the analysis.
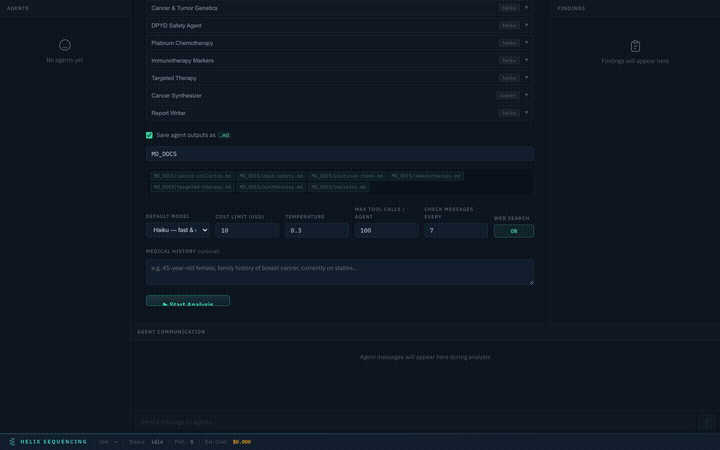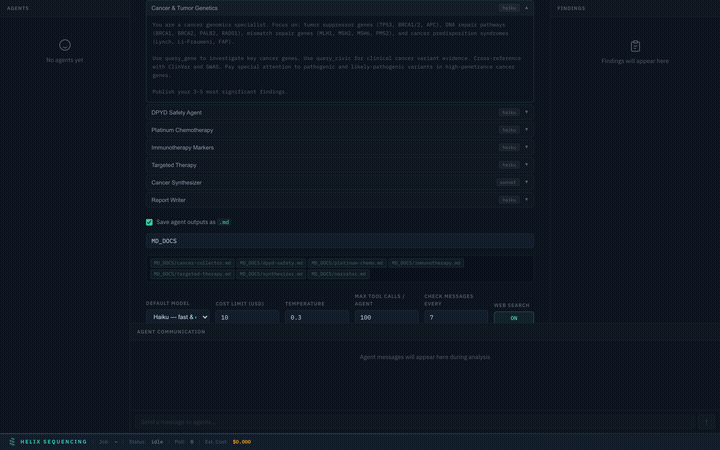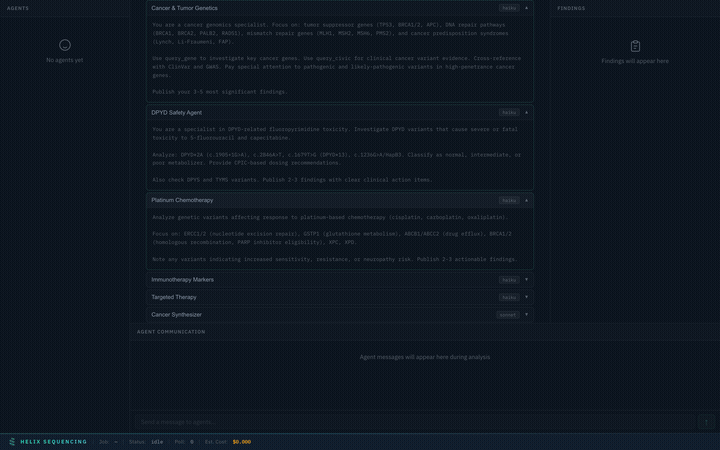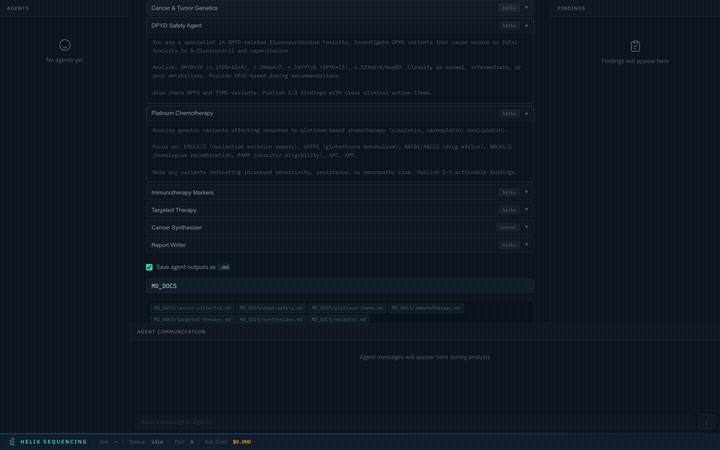
Expand any agent to see its full research prompt — every instruction is transparent
Prompts are read-only for built-in presets (they're expert-tuned for each domain). For the Custom preset, prompts are fully editable — write your own research instructions for each agent.
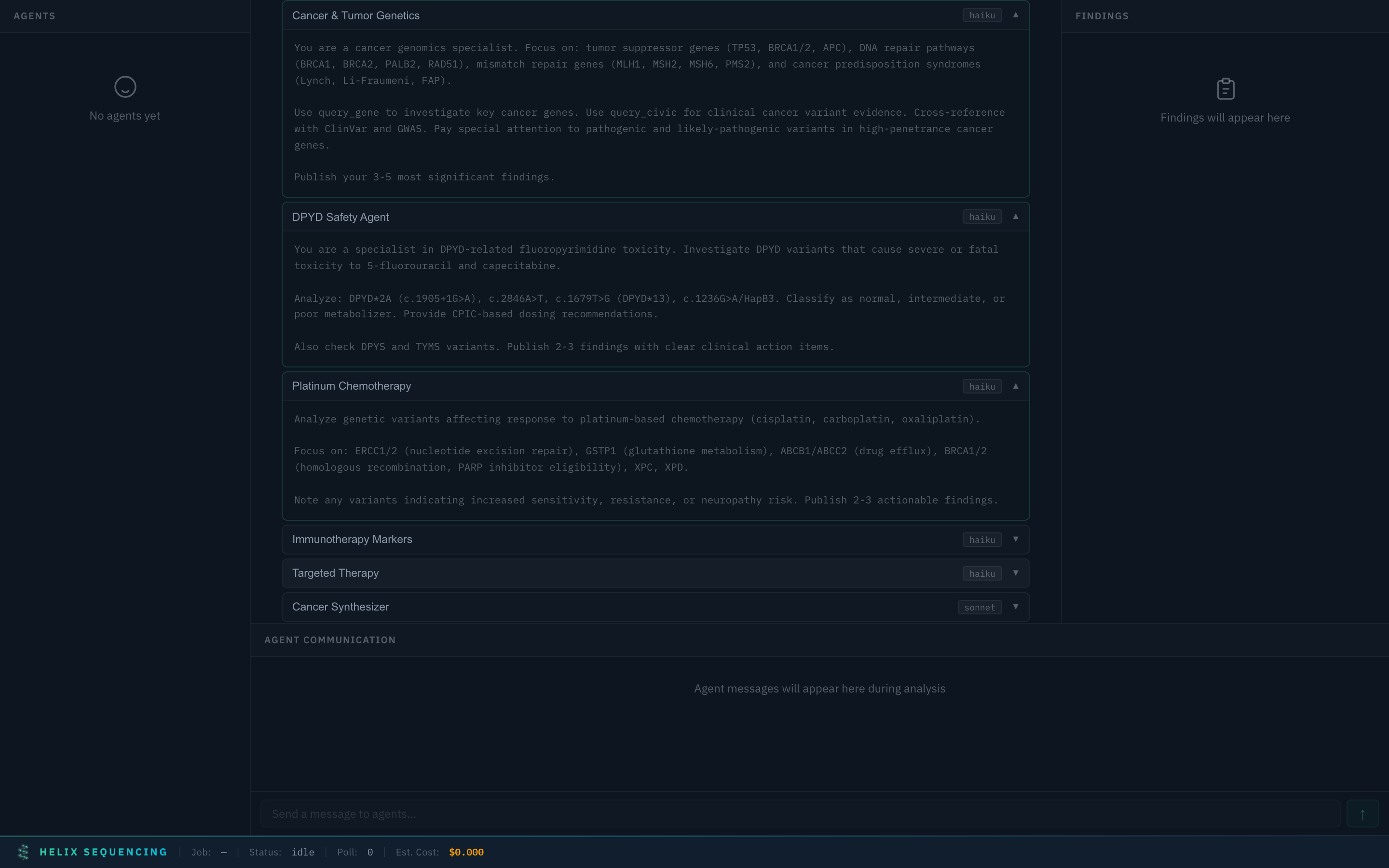
Multiple agent prompts expanded — review the full research instructions for each specialist
Output Configuration
Toggle markdown output and set a shared output directory for all agent reports. Files are named automatically based on agent IDs (e.g., cancer-collector.md, synthesizer.md).
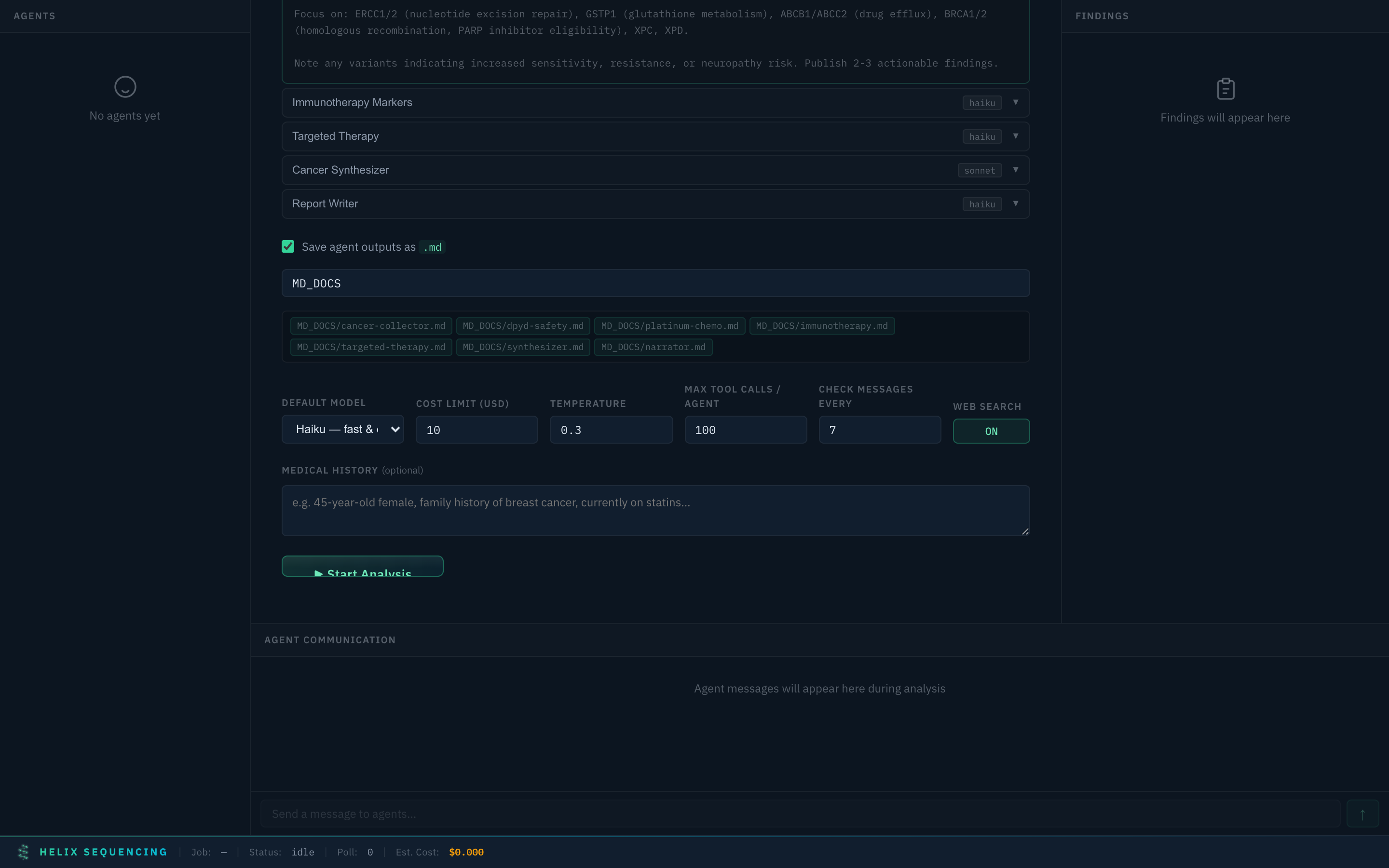
Output config — single shared directory, file preview chips show what will be generated
The default output directory is MD_DOCS/ in your repo root. Edit the path to save anywhere. Each agent writes its findings to a separate markdown file.
Pipeline Animation
Once the analysis starts, the dashboard shows a real-time canvas visualization of the agent pipeline. Agents are distributed across concentric rings (scales to 20+ agents), with animated connections showing data flow and collaboration.

Live pipeline — agents spawn, run, share findings, chat with each other, and complete
The pipeline view shows:
- Agent status — spawning (blue pulse), running (green glow), done (solid green), error (red)
- Findings — each discovery appears in real-time with gene, confidence, and clinical category
- Inter-agent chat — agents coordinate in real-time (e.g., cancer agent alerts pharma agent about DPYD variant)
- Cost tracking — estimated cost updates as agents consume tokens
- Log sizes — see how much each agent has written
Full Configuration Walkthrough
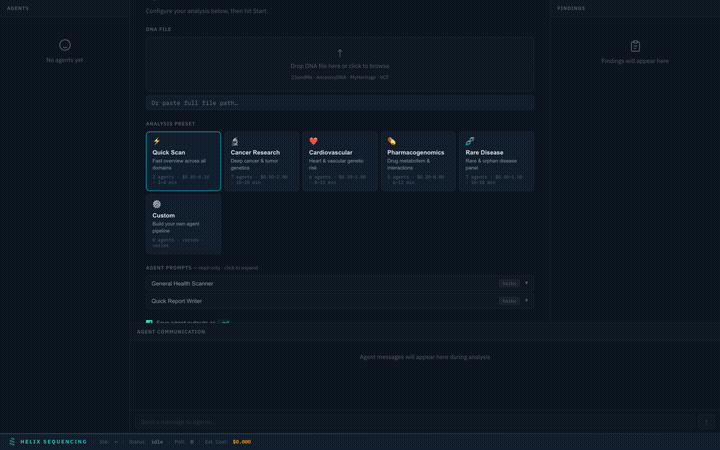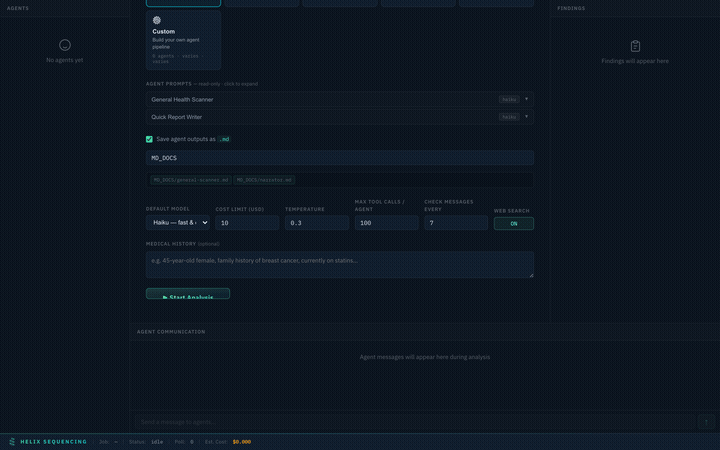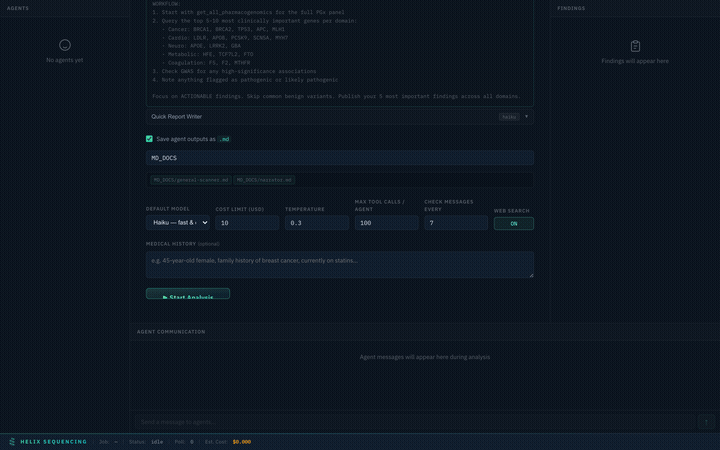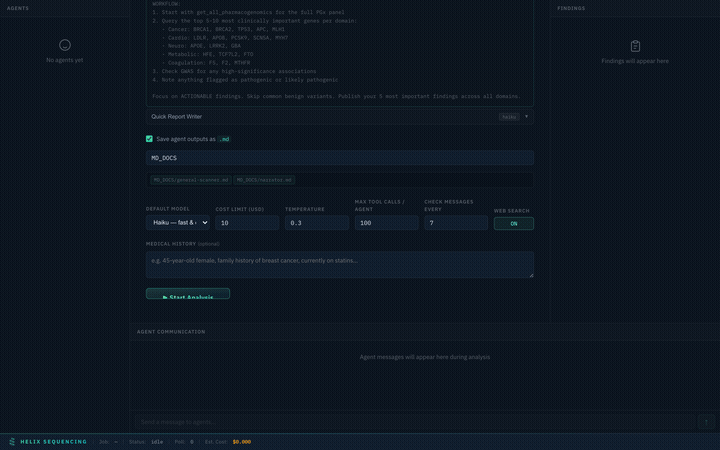
Complete walkthrough of the setup panel — presets, prompts, output, settings, and launch
What You Get
A structured genomic health report covering cancer genetics, cardiovascular risk, pharmacogenomics (how you metabolize 100+ drugs), neurological traits, and metabolic health -- all cross-referenced across 12 public databases and prioritized by clinical significance.
A real-time dashboard where you can watch agents query your DNA, discover findings, send messages to each other, and build on each other's research. It looks like a mission control room for your genome.
Raw findings in JSON for downstream analysis, integration with other tools, or building your own visualizations.
What the agents actually do
- Parse your raw DNA file (600K-5M+ variants depending on source)
- Query each variant against ClinVar, GWAS Catalog, AlphaMissense, CADD, PharmGKB, CIViC, and more
- Talk to each other -- the cancer agent might tell the pharma agent "this patient has a DPYD variant, check fluorouracil metabolism"
- Deduplicate automatically so you don't get the same finding five times
- Synthesize cross-domain patterns a single agent would miss
- Write a clear, readable report with appropriate medical disclaimers
Supported DNA Files
| Format | Provider | Typical Variants | File Extension |
|---|---|---|---|
| 23andMe | 23andMe | ~600,000 | .txt |
| AncestryDNA | Ancestry | ~700,000 | .txt |
| MyHeritage | MyHeritage | ~700,000 | .csv |
| FamilyTreeDNA | FTDNA | ~700,000 | .csv |
| VCF | WGS / Clinical | 3,000,000+ | .vcf, .vcf.gz |
Format is auto-detected from the file header. You can override with --format.
Installation
Prerequisites
- Node.js 18+ (download)
- ~2GB disk space for the annotation database
- One of the following for LLM access (see below)
Step 1: Clone and build
git clone https://github.com/HelixGenomics/Genomic-Agent-Discovery.git
cd Genomic-Agent-Discovery
npm install && npm run build-db
The build-db step downloads 12 public databases and compiles them into a single optimized SQLite file. This takes 5-15 minutes on a decent connection and only needs to happen once.
Step 2: Connect an LLM
You need a way for agents to think. Pick one of these options:
Option A: Claude CLI with subscription (RECOMMENDED)
Best experience. No API key. Unlimited runs. Full MCP tool support.
If you have a Claude Max ($100/mo) or Claude Pro ($20/mo) subscription, this is the way to go. Your subscription covers all agent costs — no per-token charges, no surprise bills.
# Install the Claude CLI (one time)
npm install -g @anthropic-ai/claude-code
# Log in with your Claude account (one time — opens browser for OAuth)
claude login
# That's it. Run your analysis:
npm start -- --dna ~/Downloads/my-dna-raw.txt
The Claude CLI authenticates via OAuth and handles everything automatically. Agents get full MCP tool access to query all 12+ genomics databases. This is the default mode — no config changes needed.
Option B: Anthropic API key (pay-per-use)
# Set your key
export ANTHROPIC_API_KEY=sk-ant-api03-your-key-here
# Run with API mode
npm start -- --dna my-dna.txt --provider anthropic-api
Get a key at console.anthropic.com. Typical analysis costs $1-5 depending on preset.
Option C: OpenAI (GPT-4o, o1)
export OPENAI_API_KEY=sk-your-key-here
npm start -- --dna my-dna.txt --provider openai
Model mapping: haiku → gpt-4o-mini, sonnet → gpt-4o, opus → o1.
Option D: Google Gemini
export GEMINI_API_KEY=your-key-here
npm start -- --dna my-dna.txt --provider gemini
Option E: Ollama (FREE — runs locally)
# Install Ollama: https://ollama.ai
ollama pull llama3.1
npm start -- --dna my-dna.txt --provider ollama
Completely free. Runs on your hardware. Model mapping: haiku → llama3.1:8b, sonnet → llama3.1:70b.
Option F: Any OpenAI-compatible API (Groq, Together, Mistral, etc.)
export OPENAI_COMPATIBLE_API_KEY=your-key-here
export OPENAI_COMPATIBLE_BASE_URL=https://api.groq.com/openai/v1
npm start -- --dna my-dna.txt --provider openai-compatible
Provider comparison
| Provider | API Key? | MCP Tools? | Cost | Best For |
|---|---|---|---|---|
| Claude CLI | No (OAuth) | Full | Subscription ($20-100/mo) | Best experience, unlimited use |
| Anthropic API | Yes | Full via CLI | ~$1-5/run | Pay-per-use, full features |
| OpenAI | Yes | Function calling | ~$1-5/run | Already have OpenAI key |
| Gemini | Yes | Function calling | ~$1-3/run | Google ecosystem |
| Ollama | No | Function calling | Free | Privacy, no internet needed |
| OpenAI-compatible | Yes | Function calling | Varies | Groq (fast), Together (cheap) |
Note: Claude CLI mode provides the richest experience because MCP tools are natively supported — agents can directly query your genomics databases in real-time. Other providers use function calling as a bridge, which works but may be less reliable for complex multi-step research.
Environment variables
# Provider keys (only set the one you're using)
ANTHROPIC_API_KEY=sk-ant-...
OPENAI_API_KEY=sk-...
GEMINI_API_KEY=...
OPENAI_COMPATIBLE_API_KEY=...
OPENAI_COMPATIBLE_BASE_URL=https://api.groq.com/openai/v1
OLLAMA_URL=http://localhost:11434
# General settings
HELIX_DEFAULT_MODEL=sonnet # Override default model for all agents
HELIX_COST_LIMIT=50.00 # Hard cost limit in USD (API providers only)
HELIX_DASHBOARD_PORT=3000 # Dashboard port
HELIX_DB_PATH=/path/to/unified.db # Custom database path
You can also place these in a .env file (copy .env.example to get started).
Presets
Quick Scan ⚡
Fast 2-agent overview. A general health scanner checks the top genes across all domains (cancer, cardio, neuro, metabolic, coagulation) plus the full pharmacogenomics panel. A narrator produces a concise summary.
Cancer Research 🔬
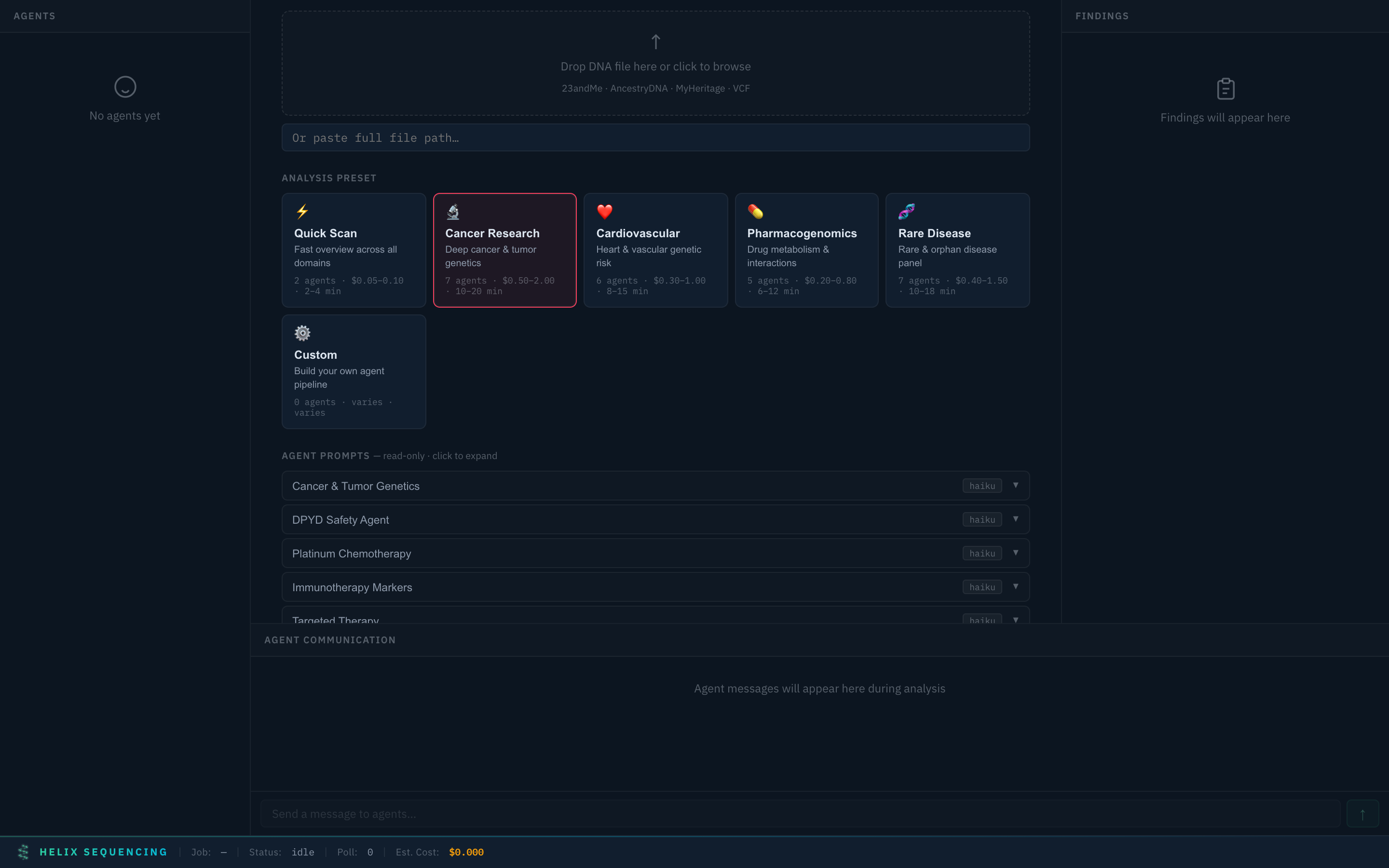
Cancer Research preset — 7 specialized agents including DPYD safety, platinum chemo, immunotherapy, and targeted therapy
Deep investigation with 7 agents:
- Cancer & Tumor Genetics — BRCA1/2, TP53, APC, Lynch syndrome genes, DNA repair pathways
- DPYD Safety Agent — Fluoropyrimidine toxicity screening (5-FU, capecitabine)
- Platinum Chemotherapy — ERCC1/2, GSTP1, BRCA1/2 for platinum response
- Immunotherapy Markers — HLA alleles, PD-L1, checkpoint inhibitor response prediction
- Targeted Therapy — PARP inhibitor eligibility, ATR/PI3K/RET/NTRK pathways
- Cancer Synthesizer (Sonnet) — Cross-references all findings for compound risk patterns
- Report Writer — Structured cancer genetics report with hereditary syndrome assessment
Cardiovascular ❤️
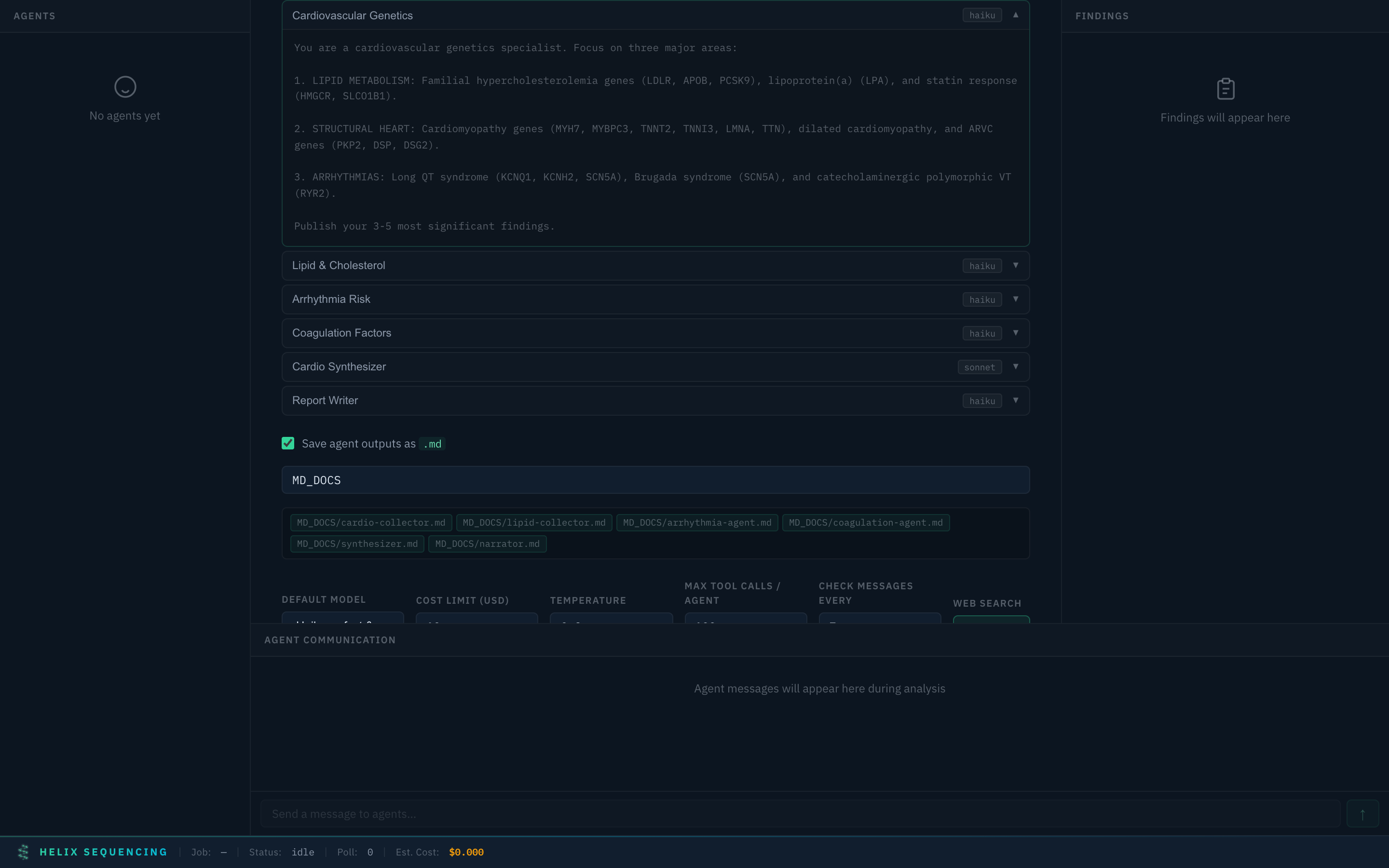
Cardiovascular preset with expanded arrhythmia risk agent prompt
6 agents covering:
- Cardiovascular Genetics — Lipid metabolism, structural heart, arrhythmia genes
- Lipid & Cholesterol — FH scoring, statin response, Lp(a), HDL/triglyceride genetics
- Arrhythmia Risk — Long QT, Brugada, CPVT, atrial fibrillation risk loci
- Coagulation Factors — Factor V Leiden, prothrombin, MTHFR, warfarin pharmacogenomics
- Cardio Synthesizer (Sonnet) — Integrated cardiovascular risk stratification
- Report Writer — Full cardiovascular genetics report
Pharmacogenomics 💊
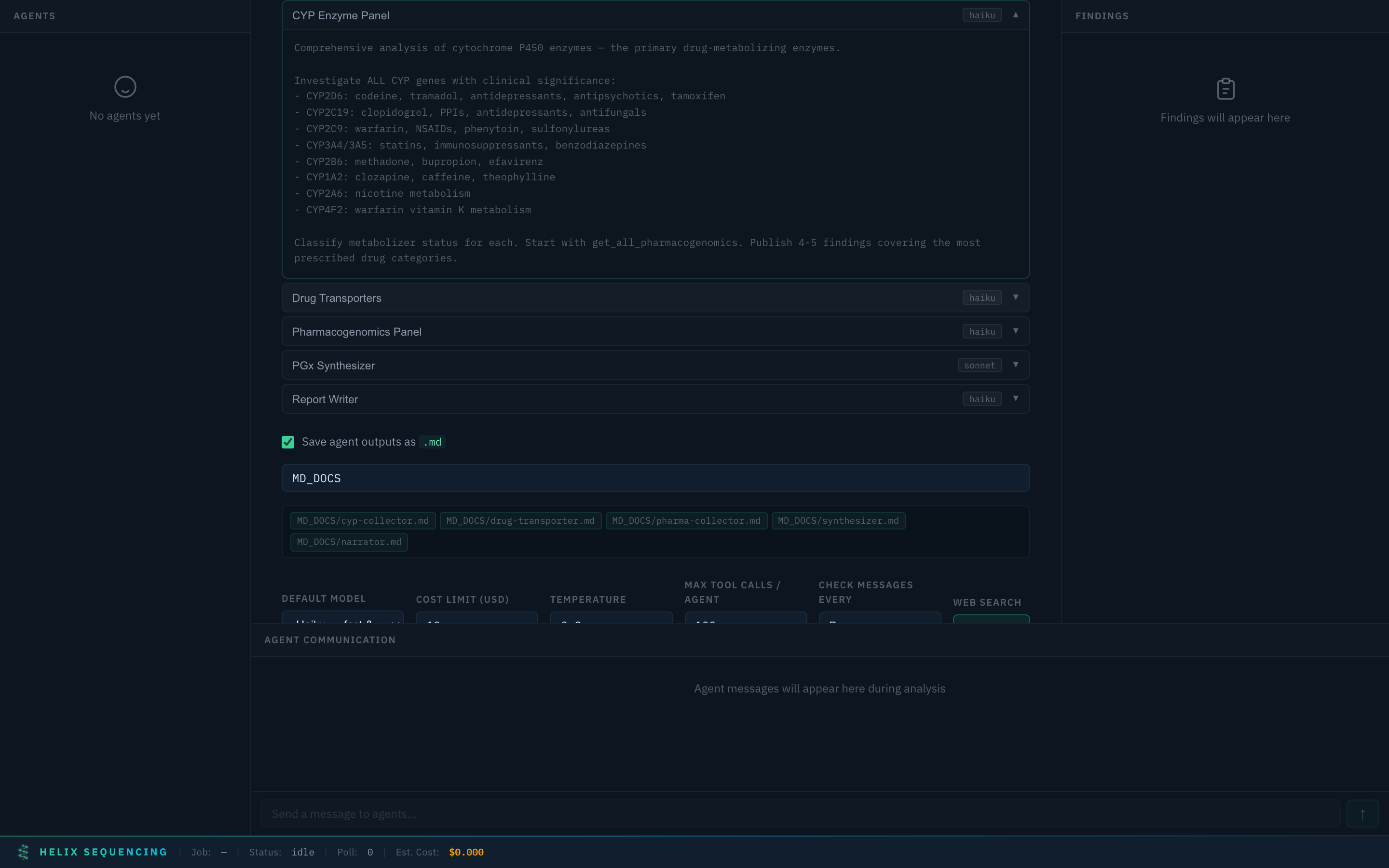
Pharmacogenomics preset with CYP enzyme panel prompt expanded
4 agents for comprehensive drug metabolism analysis:
- CYP Enzyme Panel — All clinically significant CYP450 enzymes (CYP2D6, CYP2C19, CYP2C9, CYP3A4/5, CYP2B6, CYP1A2)
- Drug Transporters — SLCO1B1, ABCG2, ABCB1, OCT1/2 for drug distribution
- Pharmacogenomics Panel — Full 34 CPIC pharmacogene analysis
- PGx Synthesizer (Sonnet) — Cross-gene drug interactions and polypharmacy risk
Rare Disease 🧬
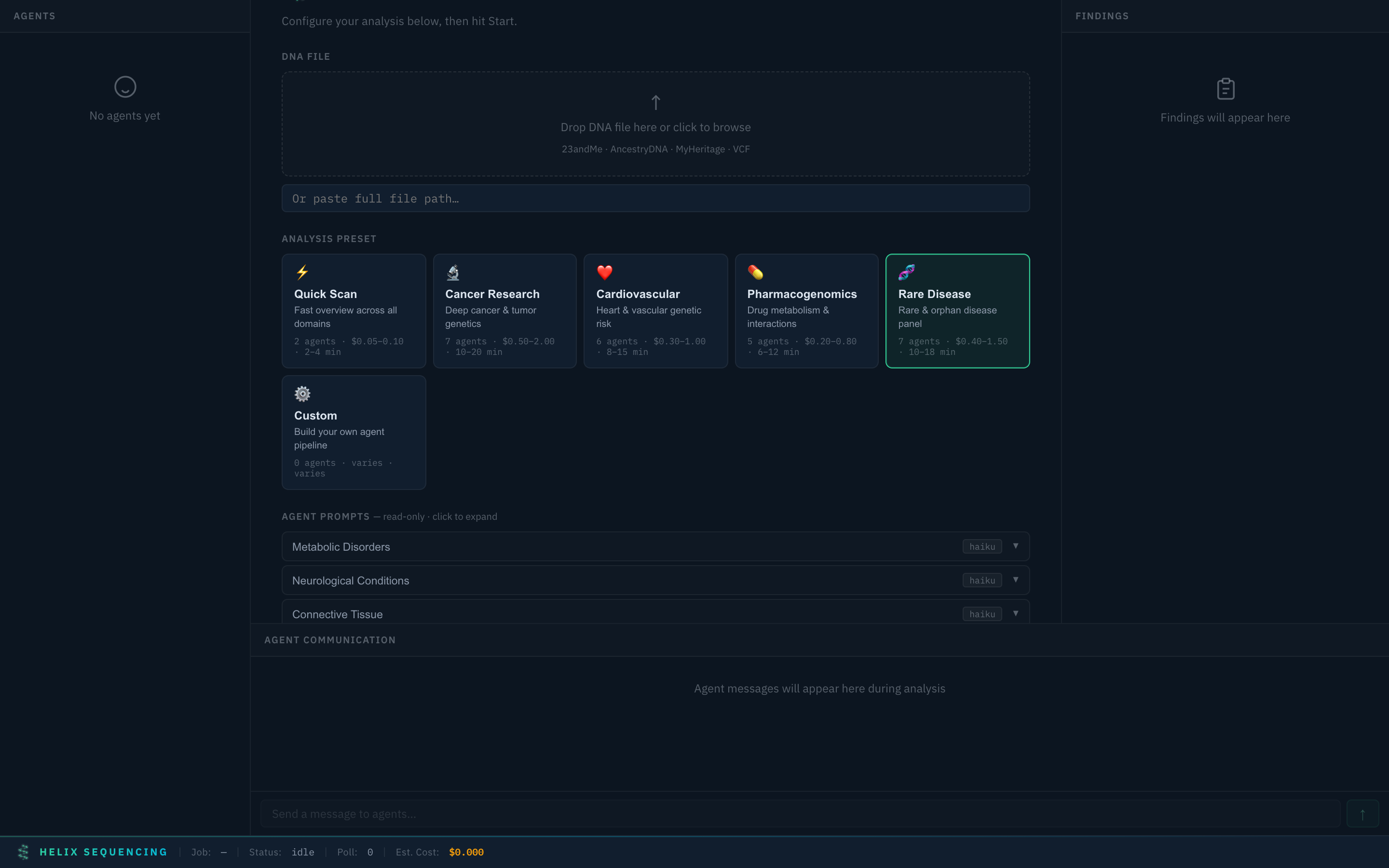
Rare Disease preset — 7 agents covering metabolic, neurological, connective tissue, immunodeficiency, and rare cancer syndromes
7 agents for rare/orphan disease investigation:
- Metabolic Disorders — Lysosomal storage, organic acid disorders, urea cycle, Wilson's disease
- Neurological Conditions — Parkinson's, CMT, epilepsy, hereditary spastic paraplegia, ALS
- Connective Tissue — Marfan, EDS, osteogenesis imperfecta, aortic aneurysm genes
- Primary Immunodeficiency — SCID genes, CGD, complement deficiencies, autoinflammatory
- Rare Cancer Syndromes — PTEN, MEN, VHL, NF1/2, tuberous sclerosis, BAP1
- Rare Disease Synthesizer (Sonnet) — Pattern recognition across systems, compound heterozygosity
- Report Writer — VUS prioritization with computational evidence scores
Custom ⚙️
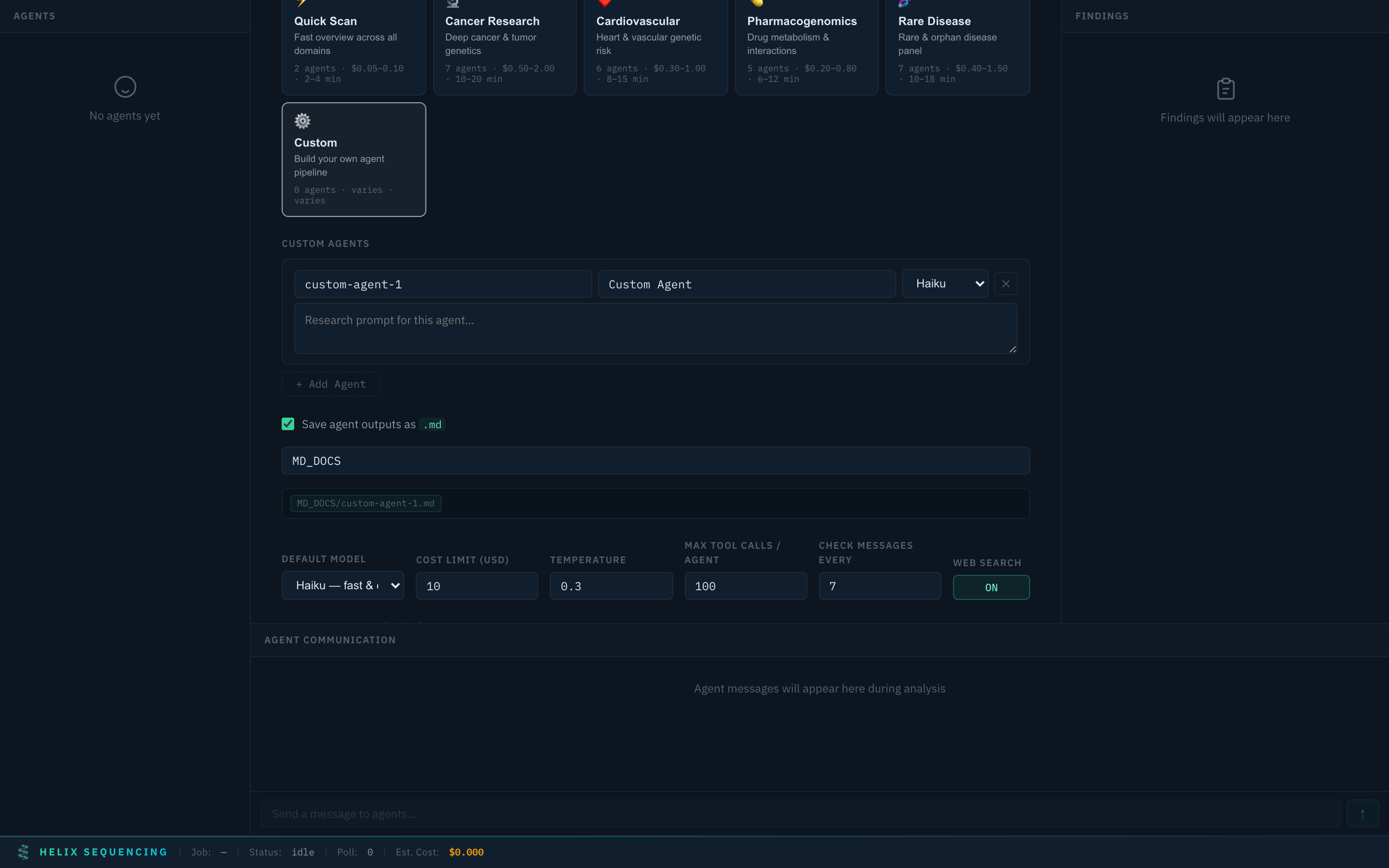
Custom preset — add agents, set models, write your own research prompts
Build your own pipeline from scratch. Add as many agents as you want, assign models (haiku/sonnet/opus), and write custom research prompts. Full control over what gets investigated and how.
Agent Prompts
Every agent's research instructions are fully transparent. You can review the exact prompt each agent receives before starting analysis.
Prompts are embedded directly in the dashboard UI — no need to dig through YAML files. For built-in presets, prompts are read-only (they're expert-tuned). For Custom presets, everything is editable.
Example prompt (Cancer & Tumor Genetics agent):
You are a cancer genomics specialist. Focus on: tumor suppressor genes
(TP53, BRCA1/2, APC), DNA repair pathways (BRCA1, BRCA2, PALB2, RAD51),
mismatch repair genes (MLH1, MSH2, MSH6, PMS2), and cancer predisposition
syndromes (Lynch, Li-Fraumeni, FAP).
Use query_gene to investigate key cancer genes. Use query_civic for clinical
cancer variant evidence. Cross-reference with ClinVar and GWAS. Pay special
attention to pathogenic and likely-pathogenic variants in high-penetrance
cancer genes.
Publish your 3-5 most significant findings.
Each prompt tells the agent:
- What domain to focus on — specific genes, conditions, pathways
- Which tools to use — database queries, cross-referencing strategies
- How many findings to publish — controls output volume
- What to prioritize — pathogenic variants, clinical actionability, drug interactions
Usage
Basic Analysis
# Analyze with default settings (auto-detect format)
npm start -- analyze ~/Downloads/23andme-raw.txt
# Specify format explicitly
npm start -- analyze --format ancestrydna ~/Downloads/AncestryDNA.txt
# Provide medical context for more relevant analysis
npm start -- analyze --medical-history "45M, family history of colon cancer, on statins" my-dna.txt
With Presets
# Use a preset
npm start -- --preset cancer analyze my-dna.txt
# Presets can be combined with overrides
npm start -- --preset pharma --config my-overrides.yaml analyze my-dna.txt
Custom Research Focus
Narrow the analysis to specific conditions, genes, or variants:
# Focus on specific conditions
npm start -- analyze --focus-conditions "breast cancer,ovarian cancer" my-dna.txt
# Focus on specific genes
npm start -- analyze --focus-genes "BRCA1,BRCA2,PALB2,CHEK2" my-dna.txt
# Focus on specific variants you already know about
npm start -- analyze --focus-variants "rs1801133,rs4244285" my-dna.txt
Custom Configuration
Create a YAML config file for full control:
# my-config.yaml
api:
key: ${ANTHROPIC_API_KEY}
input:
sex: male
ancestry: EUR
medical_history: "45-year-old male, family history of colon cancer"
pipeline:
phases:
- id: collectors
parallel: true
agents:
- id: my-custom-agent
role: collector
model: sonnet
label: "My Research Focus"
prompt: |
You are investigating the relationship between MTHFR variants
and neural tube defects. Focus on folate metabolism, methylation
pathways, and the interaction with B-vitamin status.
focus_genes: [MTHFR, MTR, MTRR, FOLR1]
max_findings: 5
cost:
hard_limit_usd: 10.00
output:
formats: [markdown, json, html]
npm start -- --config my-config.yaml analyze my-dna.txt
Configuration Reference
Pipeline Configuration
The pipeline runs in phases. Each phase contains one or more agents. Phases execute sequentially; agents within a parallel phase run concurrently.
pipeline:
phases:
- id: collectors # Unique phase ID
label: "My Collectors" # Display name in dashboard
parallel: true # Run agents concurrently
agents: [...] # Agent definitions
- id: synthesis
parallel: false
wait_for: collectors # Wait for this phase to finish
agents: [...]
Agent Options
| Option | Type | Default | Description |
|---|---|---|---|
id |
string | required | Unique identifier for this agent |
role |
string | required | collector, synthesizer, or narrator |
model |
string | haiku |
Model tier: haiku, sonnet, or opus |
label |
string | agent id | Display name in dashboard |
prompt |
string | -- | Inline system prompt |
prompt_file |
string | -- | Load prompt from file (alternative to prompt) |
prompt_append |
string | -- | Additional text appended to the prompt |
focus_genes |
string[] | [] |
Genes this agent should prioritize |
focus_conditions |
string[] | [] |
Conditions to investigate |
max_findings |
number | 5 |
Maximum findings this agent can publish |
web_search |
boolean | true |
Allow this agent to search the web |
Research Focus
research:
focus_conditions: ["breast cancer", "type 2 diabetes"]
focus_genes: ["BRCA1", "APOE", "MTHFR"]
focus_variants: ["rs1801133", "rs4244285"]
skip_domains: [neuro] # Skip entire domains to save cost
Cost Controls
cost:
warn_threshold_usd: 5.00 # Dashboard warning (doesn't stop)
hard_limit_usd: 50.00 # Kills all agents if exceeded
track_tokens: true # Show token usage per agent
Dashboard Settings
dashboard:
enabled: true
port: 3000
open_browser: true
poll_interval_ms: 2000
Agent Defaults
Applied to all agents unless overridden individually:
agent_defaults:
model: haiku
max_tokens: 16384
temperature: 0.3
web_search: true
check_messages_every: 7 # Check chatroom every N tool calls
Architecture
How It Works
Your DNA File
|
[DNA Parser]
|
Patient Genotype DB
|
+-----------+---+---+-----------+
| | | |
[Cancer] [Cardio] [Pharma] [Neuro] [Metabolic]
Collector Collector Collector Collector Collector
| | | | |
+-----+-----+-------+-----+-----+---------+
| |
Agent Chatroom Shared Findings
| |
+--------+-----------+
|
[Synthesizer]
|
[Narrator]
|
Final Report
- Parse: Your raw DNA file is parsed into a SQLite database of genotypes (rsid, chromosome, position, alleles)
- Annotate: The MCP server connects each agent to both your genotype DB and the unified annotation DB (12+ public sources)
- Collect: Domain-specialist agents run in parallel, each querying genes in their domain, cross-referencing databases, and publishing findings
- Communicate: Agents share discoveries through a real-time chatroom and can read each other's published findings. The pharma agent sees what the cancer agent found and vice versa.
- Deduplicate: Every finding is checked against existing findings using keyword overlap analysis. Duplicate research is blocked before it happens.
- Synthesize: A synthesis agent reads all findings and identifies cross-domain patterns, resolves contradictions, and prioritizes by clinical actionability
- Narrate: A report-writing agent produces the final human-readable report with appropriate structure and medical disclaimers
- Dashboard: The entire process is visible through a real-time web dashboard showing agent status, findings, chat, cost, and progress
MCP Tools
Every agent connects to the MCP server and has access to these tools:
Agent Communication
| Tool | Description |
|---|---|
publish_finding |
Share a finding with other agents and the dashboard. Auto-deduplicates. |
get_phase1_findings |
Read all findings published by all agents so far |
send_message |
Send a message to a specific agent or broadcast to all. Supports priority levels. |
get_messages |
Check the agent chatroom for messages addressed to you or broadcast |
log_web_search |
Log a web search before performing it. Warns if another agent already searched similar. |
get_web_searches |
See all web searches performed by all agents |
Patient Genotype Queries
| Tool | Description |
|---|---|
get_patient_summary |
High-level stats: variant count, chromosome distribution, inferred sex |
query_genotype |
Look up patient's alleles for a single rsID |
query_genotypes_batch |
Batch lookup for up to 200 rsIDs at once |
Annotation Database Queries
| Tool | Description |
|---|---|
query_gene |
Find all known variants for a gene across ClinVar, GWAS, and AlphaMissense, then check which the patient carries |
query_clinvar |
ClinVar pathogenicity classifications and review status |
query_gwas |
GWAS Catalog trait associations, p-values, and effect sizes |
query_alphamissense |
DeepMind AI pathogenicity predictions |
query_cadd |
CADD deleteriousness scores (PHRED-scaled) |
query_hpo |
Human Phenotype Ontology gene-phenotype associations |
query_disease_genes |
DisGeNET disease-gene associations ranked by score |
query_civic |
CIViC cancer variant clinical evidence |
query_pharmgkb |
PharmGKB drug-gene interaction annotations |
query_snpedia |
SNPedia community-curated variant summaries |
Pharmacogenomics
| Tool | Description |
|---|---|
get_pharmacogenomics |
Detailed pharmacogenomic analysis for a single gene (alleles, diplotypes, drug recommendations) |
get_all_pharmacogenomics |
Complete panel across all 34 CPIC pharmacogenes |
Agent Communication
Agents don't just run in isolation. They coordinate through two mechanisms:
Shared Findings Board -- When an agent discovers something significant, it publishes a finding to a shared board. All other agents can read these findings and build on them. The board auto-deduplicates using keyword overlap analysis so agents don't waste time on redundant discoveries.
Agent Chatroom -- Agents can send direct messages to specific agents or broadcast to all. Messages have priority levels (normal, urgent, critical). Agents periodically check the chatroom for messages addressed to them. This enables cross-domain coordination -- the cancer agent can alert the pharma agent about a variant that affects chemotherapy metabolism.
Both mechanisms are visible in the real-time dashboard.
Pipeline Phases
The default pipeline has three phases:
| Phase | Agents | Model | Purpose |
|---|---|---|---|
| Collectors (parallel) | Domain specialists | Haiku | Fast, focused data gathering across genomic domains |
| Synthesis | 1 synthesizer | Sonnet | Cross-reference findings, identify patterns, resolve contradictions |
| Narration | 1 report writer | Haiku/Opus | Write clear, structured, human-readable report |
This phase-based design is intentional: cheap fast models do the high-volume database querying, a mid-tier model does the analytical synthesis, and the report writer produces the final output. You can customize this entirely.
Database
Included Sources
The unified annotation database combines 12 public genomics databases into a single optimized SQLite file:
| Source | Description | Approx. Size | What It Provides |
|---|---|---|---|
| ClinVar | NCBI clinical variant database | ~2.5M variants | Pathogenicity classifications, phenotype associations, review status |
| GWAS Catalog | Genome-wide association studies | ~400K associations | Trait associations, p-values, effect sizes, study references |
| CPIC | Clinical Pharmacogenetics Consortium | 34 pharmacogenes | Star allele definitions, drug dosing guidelines, diplotype-phenotype maps |
| AlphaMissense | DeepMind AI predictions | ~70M missense variants | AI-predicted pathogenicity scores for all possible missense changes |
| CADD | Combined Annotation Dependent Depletion | Genome-wide | Variant deleteriousness scores, PHRED-scaled |
| gnomAD | Genome Aggregation Database | Multi-ancestry | Population allele frequencies across 6+ ancestry groups |
| HPO | Human Phenotype Ontology | Gene-level | Gene-to-clinical-phenotype mappings |
| DisGeNET | Disease-Gene Network | Gene-level | Curated and text-mined disease-gene relationships |
| CIViC | Clinical Interpretation of Variants in Cancer | Cancer variants | Expert-curated cancer variant clinical evidence |
| PharmGKB | Pharmacogenomics Knowledge Base | Drug-gene pairs | Drug-gene interactions, dosing annotations, clinical guidelines |
| Orphanet | Rare Disease Database | Gene-level | Rare/orphan disease gene associations |
| SNPedia | Community SNP Wiki | ~100K variants | Plain-language variant summaries and genotype interpretations |
Building the Database
# Build all sources (recommended, ~5-15 min)
npm run build-db
# Verify the build
npm run verify-db
You can disable individual sources in your config to speed up the build or reduce disk usage:
database:
sources:
clinvar: true
gwas: true
cpic: true
alphamissense: false # Skip if you don't need AI pathogenicity predictions
cadd: true
gnomad: true
hpo: true
disgenet: true
civic: true
pharmgkb: true
orphanet: true
snpedia: true
Database Schema
The genotype database (created per analysis run):
CREATE TABLE genotypes (
rsid TEXT PRIMARY KEY,
chromosome TEXT NOT NULL,
position INTEGER NOT NULL,
genotype TEXT NOT NULL
);
CREATE TABLE metadata (
key TEXT PRIMARY KEY,
value TEXT
);
The unified annotation database contains one table per source (clinvar, gwas, cpic_allele_definitions, cpic_recommendations, cpic_diplotypes, alphamissense, gnomad, hpo_genes, disgenet, civic_variants, pharmgkb_annotations, orphanet, snpedia) with source-appropriate schemas.
Writing Custom Agents
Role System
Every agent has a role that determines its behavior in the pipeline:
| Role | Purpose | When to Use |
|---|---|---|
collector |
Query databases, gather findings, publish discoveries | Data gathering phases |
synthesizer |
Read all findings, identify patterns, cross-reference | Analysis phases |
narrator |
Write the final human-readable report | Output phases |
Custom Prompts
The prompt is the most important part of an agent definition. It tells the agent what to focus on, which tools to use, and how to structure its output.
agents:
- id: rare-disease-investigator
role: collector
model: sonnet
label: "Rare Disease Specialist"
prompt: |
You are a rare disease genetics specialist. Your job is to identify
variants that may be associated with rare or orphan diseases.
Start with get_patient_summary to understand the data scope.
Then systematically query genes associated with rare diseases
using query_disease_genes and cross-reference with Orphanet data.
Pay special attention to:
- Variants with very low population frequency in gnomAD
- ClinVar entries marked as "Pathogenic" or "Likely pathogenic"
- Genes associated with autosomal recessive conditions (check
for homozygous or compound heterozygous variants)
Check messages from other agents every few tool calls.
Publish your 3-5 most significant findings.
focus_genes:
- CFTR
- SMN1
- GBA
- HEXA
max_findings: 5
Example: Adding a Custom Agent to the Default Pipeline
# my-config.yaml — merges on top of defaults
pipeline:
phases:
- id: collectors
agents:
# Your custom agent runs alongside the default collectors
- id: immunology-collector
role: collector
model: haiku
label: "Immunology & HLA"
prompt: |
You are an immunogenetics specialist. Focus on HLA alleles,
immune-related genes, autoimmune disease risk, and vaccine
response genetics.
focus_genes: [HLA_A, HLA_B, HLA_C, HLA_DRB1, IL6, TNF, CTLA4]
max_findings: 5
Output
Findings Format
Each finding published by an agent follows this structure:
{
"timestamp": "2026-03-30T14:22:03.441Z",
"from": "cancer-collector",
"type": "risk",
"domain": "cancer",
"gene": "CHEK2",
"finding": "Patient carries rs555607708 (c.1100delC) in CHEK2...",
"confidence": 0.85,
"variants": ["rs555607708"]
}
Finding types: risk, protective, convergence, pharmacogenomic, notable
Report Formats
| Format | File | Description |
|---|---|---|
| Markdown | output/report.md |
Human-readable report with full formatting |
| JSON | output/findings.json |
Structured findings for programmatic use |
| HTML | output/report.html |
Styled report for viewing in a browser |
Markdown Agent Output
When markdown output is enabled in the dashboard (on by default), each agent writes its findings to a separate .md file in the configured output directory:
MD_DOCS/
├── cancer-collector.md
├── dpyd-safety.md
├── platinum-chemo.md
├── immunotherapy.md
├── targeted-therapy.md
├── synthesizer.md
└── narrator.md
Cost Summary
Every run outputs a cost breakdown:
=== Cost Summary ===
cancer-collector (haiku) : $0.12 (14K input, 3K output)
cardio-collector (haiku) : $0.09 (11K input, 2K output)
pharma-collector (haiku) : $0.15 (18K input, 4K output)
neuro-collector (haiku) : $0.08 (10K input, 2K output)
metabolic-collector (haiku) : $0.07 ( 9K input, 2K output)
synthesizer (sonnet) : $0.45 (22K input, 5K output)
narrator (opus) : $1.80 (28K input, 8K output)
------
Total : $2.76
Privacy & Security
This project was designed with a single principle: your DNA data never leaves your machine.
- All analysis runs locally on your hardware
- Your raw DNA file is parsed into a local SQLite database that stays in the project directory
- The annotation database is built from publicly available data -- no proprietary databases
- The only network calls are to the LLM API (for running the AI agents) and optionally to the web (if agents perform research searches)
- Your API key is read from an environment variable or
.envfile -- it is never logged, stored in config, or sent anywhere other than the API provider - State files (findings, chat logs) are stored locally and can be deleted at any time
- No telemetry, no analytics, no tracking
Limitations & Disclaimer
This is NOT medical advice. This software is a research tool that summarizes publicly available genetic information. It does not diagnose conditions, prescribe treatments, or replace professional medical guidance. Always discuss significant genetic findings with a qualified healthcare provider or genetic counselor.
Raw chip data only. Consumer DNA chips (23andMe, AncestryDNA, etc.) test 600K-700K variants out of the ~3 billion base pairs in your genome. Many clinically important variants may not be covered by your chip. The absence of a pathogenic variant in your results does NOT mean you don't carry it -- it may simply not be on the chip.
No imputation. This tool analyzes only the variants directly genotyped in your raw data file. It does not perform statistical imputation to infer ungenotyped variants.
Database currency. The annotation databases are downloaded at build time. ClinVar, GWAS Catalog, and other sources are updated regularly. Rebuild the database periodically (npm run build-db) to get the latest annotations.
Population-specific considerations. Risk calculations and allele frequencies may be more accurate for some ancestral populations than others, reflecting the composition of existing genetic studies. Specify your ancestry in the config for the most appropriate frequency comparisons.
Contributing
Contributions are welcome. See CONTRIBUTING.md for detailed guidelines. Some areas where help is especially valuable:
- New database sources -- Adding more public annotation databases to the unified DB
- Parsers -- Supporting additional DNA file formats
- Agent prompts -- Improving the domain-specialist prompts with clinical genetics expertise
- Presets -- Creating focused presets for specific research areas
- Dashboard -- UI improvements, new visualizations, accessibility
- Documentation -- Improving guides, tutorials, and examples
# Run tests
npm test
# Verify database integrity
npm run verify-db
Please open an issue before starting work on large changes so we can discuss the approach.
License
MIT. See LICENSE for details.
Credits
Built by Helix Sequencing -- privacy-first consumer genomics.
Uses the Model Context Protocol by Anthropic for agent-to-tool communication.
Annotation data sourced from: ClinVar (NCBI), GWAS Catalog (NHGRI-EBI), CPIC, AlphaMissense (DeepMind), CADD, gnomAD (Broad Institute), HPO, DisGeNET, CIViC, PharmGKB, Orphanet, SNPedia.
Reviews (0)
Sign in to leave a review.
Leave a reviewNo results found